Blog
Orbion Team
Disulfide Bond Engineering: When to Add, Remove, or Redesign

You designed a disulfide bond between residues 45 and 182 to stabilize your protein. The Cβ-Cβ distance in the AlphaFold model was 4.2 Å—perfect geometry. You expressed it in E. coli. The protein runs as a smear on non-reducing SDS-PAGE. The SEC profile shows five peaks instead of one. Your "stabilizing" disulfide created a cross-linked aggregation nightmare.
Disulfide engineering is one of the most powerful tools in protein stabilization—when it works. When it doesn't, it's one of the fastest ways to destroy a protein. The difference comes down to understanding the geometry, environment, and biology of disulfide bonds, not just the Cβ distance.
Key Takeaways
Not every cysteine pair can form a disulfide: geometry (Cβ-Cβ distance, χ3 dihedral angle), local dynamics, and redox environment all matter
E. coli cytoplasm is reducing: disulfide bonds won't form unless you use specialized strains (SHuffle, Origami) or periplasmic expression
Engineered disulfides stabilize by 2–8 kJ/mol on average, but badly placed ones destabilize by straining the backbone
Free cysteines are dangerous: unpaired Cys residues cause intermolecular disulfide-mediated aggregation—sometimes removing a disulfide is the right engineering move
Computational prediction of disulfide feasibility before mutagenesis prevents expensive failures
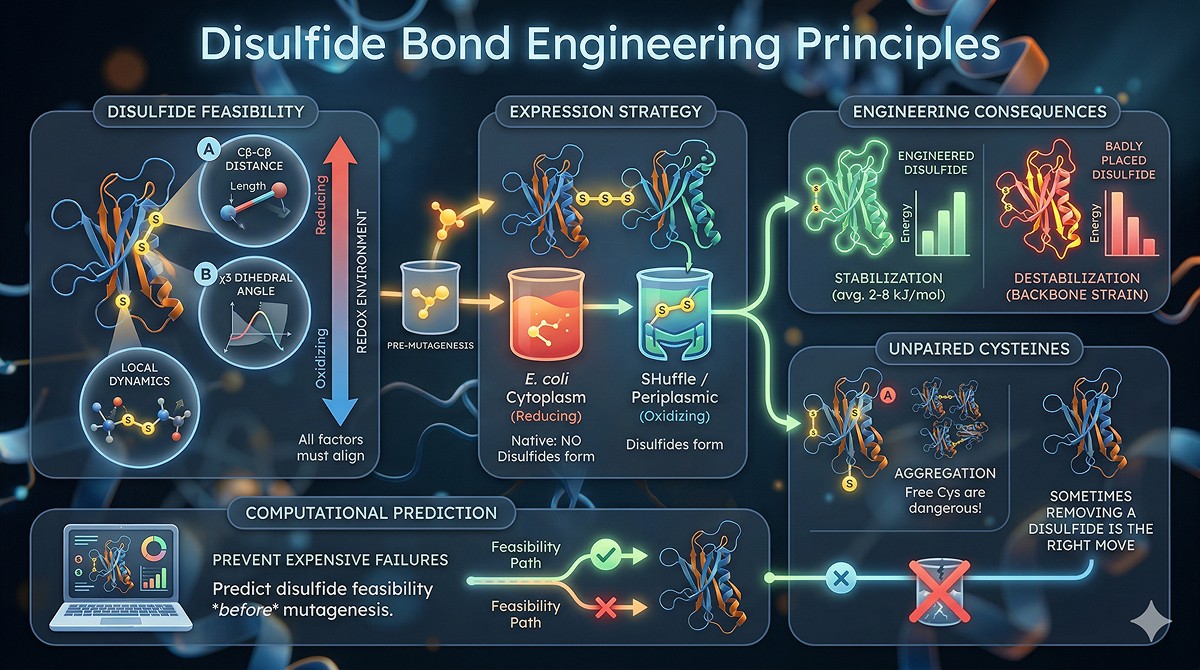
The Biology of Disulfide Bonds
What Disulfide Bonds Do
Disulfide bonds (S-S bonds between cysteine residues) serve multiple functions:
Function | Mechanism | Examples |
|---|---|---|
Structural stabilization | Cross-link distant residues, reduce conformational entropy of unfolded state | Lysozyme (4 disulfides), ribonuclease A (4 disulfides) |
Catalytic | Redox-active disulfides in enzyme mechanism | Thioredoxin, protein disulfide isomerase |
Regulatory | Redox-sensitive switches that change activity | Keap1-Nrf2, OxyR |
Assembly | Link subunits covalently | Antibody heavy-light chains, insulin A-B chains |
Where Disulfides Naturally Occur
Disulfide bonds are overwhelmingly found in secreted proteins and extracellular domains:
Only ~5% of cytoplasmic proteins have structural disulfides
The endoplasmic reticulum provides the oxidizing environment for disulfide formation
Cytoplasmic disulfides are rare because the reducing environment (glutathione, thioredoxin) keeps cysteines reduced
This has direct engineering implications: If you're adding a disulfide to a cytoplasmic protein, you're engineering something that evolution specifically avoids.
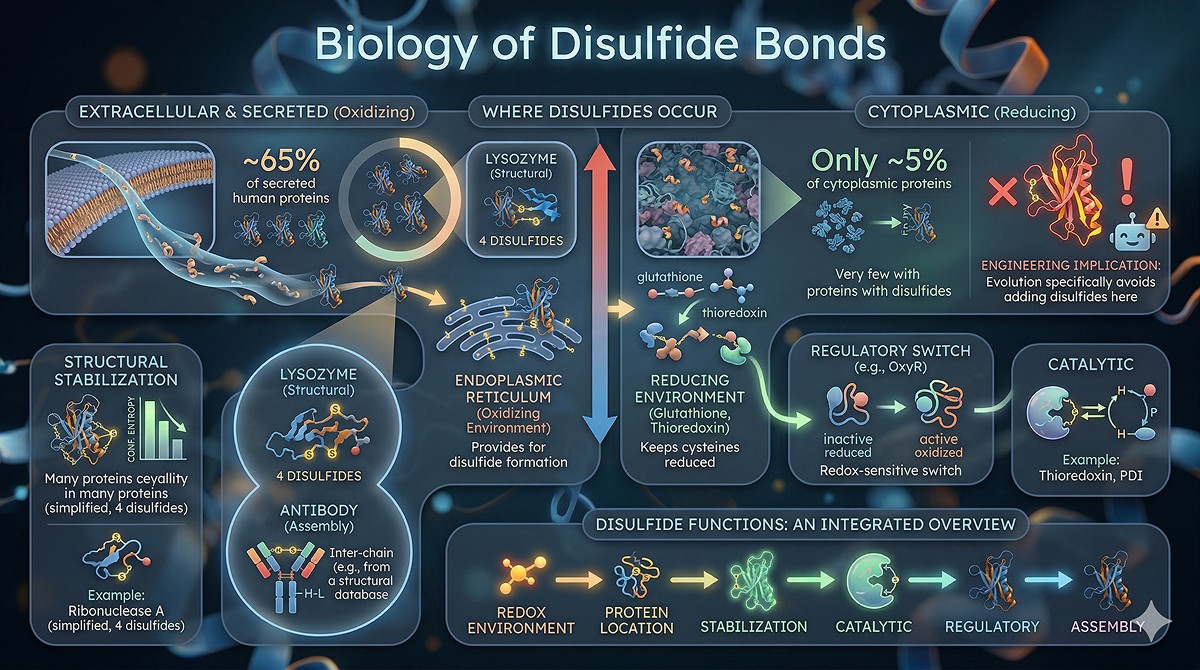
The Geometry of Disulfide Bonds
What Makes a Good Disulfide
Not all cysteine pairs form disulfides. The geometric requirements are strict (Dombkowski et al., 2014):
Parameter | Ideal Value | Acceptable Range | What Happens Outside Range |
|---|---|---|---|
Cβ-Cβ distance | 3.5–4.5 Å | 3.0–6.0 Å | Too close: backbone strain. Too far: no bond forms. |
Sγ-Sγ distance | 2.03 Å | 1.9–2.1 Å | Fixed by covalent bond length |
Cα-Cβ-Sγ angle | ~115° | 105–125° | Strain on side chain |
χ3 dihedral (Cβ-Sγ-Sγ-Cβ) | ±90° | ±60° to ±120° | Outside this range: high-energy bond |
Cα-Cα distance | 5.0–7.0 Å | 4.5–8.0 Å | Determines backbone compatibility |
The most common mistake: Using only Cβ-Cβ distance as a criterion. A pair with Cβ-Cβ = 4.0 Å but poor dihedral angles will form a strained disulfide that destabilizes the protein.
The Five Canonical Conformations
Natural disulfides adopt five well-characterized conformations based on the five dihedral angles χ1-χ2-χ3-χ2'-χ1'. The most common are:
Left-handed spiral (–, –, –, –, –): Most common, ~30% of natural disulfides
Right-handed hook (+, +, +, +, +): ~15%
Left-handed hook: ~12%
Short right-handed hook: ~10%
Various others: Remaining ~33%
Engineered disulfides that match these canonical conformations are much more likely to form correctly and stabilize the protein.
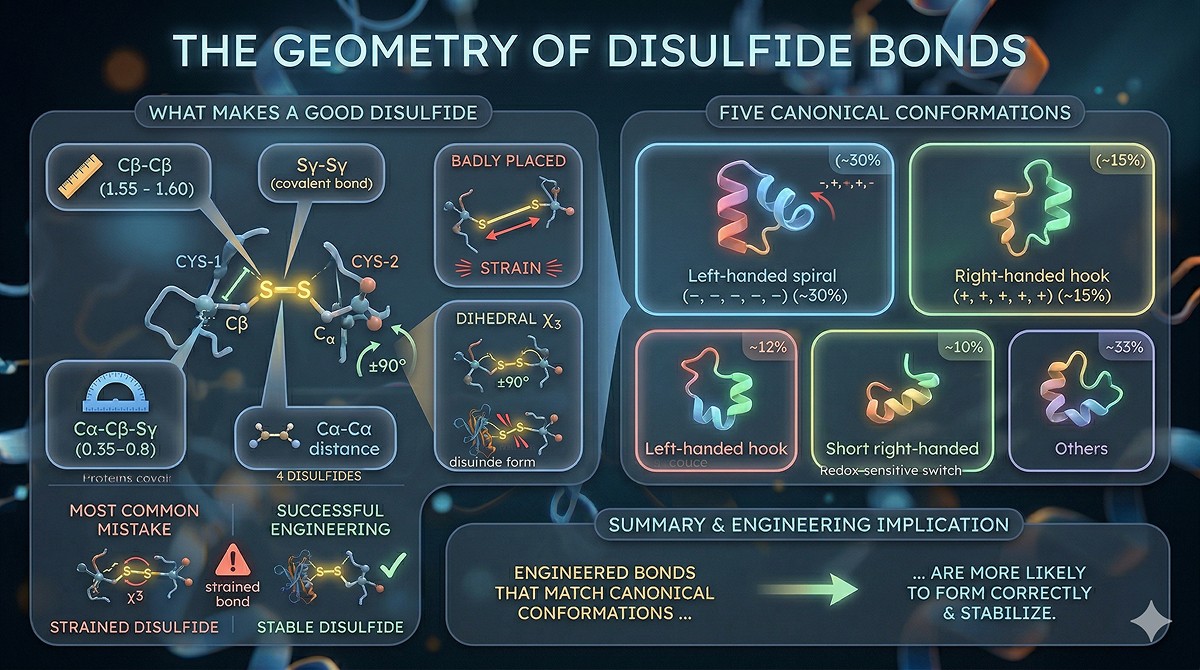
When to ADD a Disulfide Bond
The Right Targets for Disulfide Engineering
Adding a disulfide makes sense when:
You need to stabilize a protein for harsh conditions (high temperature, low pH, protease exposure)
Two residues are geometrically compatible but are naturally cysteines or can be mutated to cysteines
The protein will be in an oxidizing environment (secreted, extracellular, or expressed in a system that supports disulfide formation)
The loop or region you're stabilizing is genuinely flexible and contributes to instability
The Stabilization Mechanism
Disulfide bonds stabilize by reducing the conformational entropy of the unfolded state (Pace et al., 1988):
In the unfolded protein, residues separated by N residues in sequence have N^(3/2) possible conformations
A disulfide bond constrains these residues, reducing the entropy of the unfolded state
This increases ΔG_unfolding without significantly affecting the folded state (if geometry is good)
The entropic contribution: ΔS ≈ -2.1 - (3/2)R·ln(N) cal/mol/K, where N is the number of residues between the bonded cysteines
Practical implication: Disulfides between residues far apart in sequence stabilize more than those between nearby residues (more entropy lost upon unfolding).
Expected Stabilization
Loop Size (residues between Cys) | Expected ΔΔG | ΔTm |
|---|---|---|
10 | ~2–4 kJ/mol | +2–5°C |
25 | ~4–6 kJ/mol | +3–7°C |
50 | ~5–8 kJ/mol | +4–9°C |
100+ | ~6–10 kJ/mol | +5–12°C |
Caveat: These are theoretical maxima for well-placed disulfides. Strain in the folded state reduces the actual stabilization. Many engineered disulfides achieve only 50–70% of the theoretical maximum.
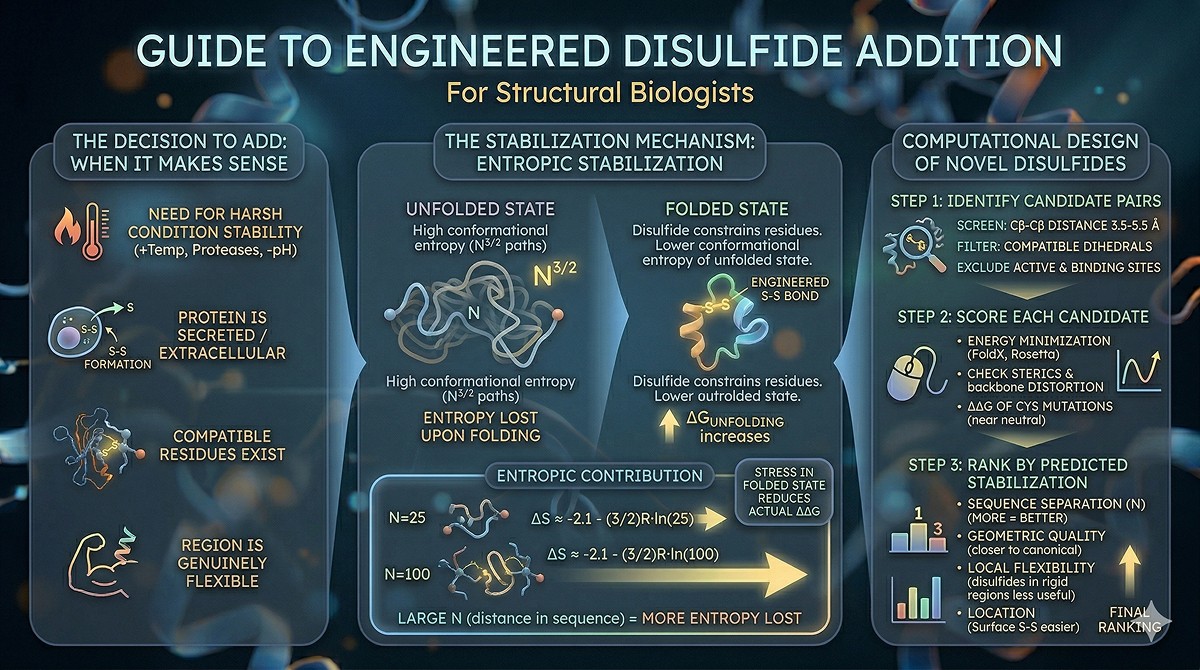
Computational Design of Novel Disulfides
Step 1: Identify candidate pairs
Screen all residue pairs for Cβ-Cβ distance 3.5–5.5 Å
Filter for compatible dihedral angles
Exclude active site residues, binding sites, PTM sites
Step 2: Score each candidate
Energy minimization after Cys mutation (Rosetta, FoldX)
Check for steric clashes
Verify that backbone doesn't distort
ΔΔG of the Cys mutations themselves (both should be near-neutral)
Step 3: Rank by predicted stabilization
Sequence separation (more = better)
Geometric quality (closer to canonical = better)
Local flexibility (disulfides in rigid regions add less value)
Location (surface vs buried—surface disulfides are easier to engineer)
When to REMOVE a Disulfide Bond
Free Cysteines: The Aggregation Problem
Sometimes the right engineering move is removing cysteines, not adding them.
The problem with free (unpaired) cysteines:
Free Cys residues can form intermolecular disulfides with Cys residues on other protein molecules → covalent aggregation
This is the most common cause of disulfide-mediated aggregation during expression, purification, and storage
Even at low concentrations, free Cys residues slowly oxidize and cross-link over days at 4°C
How common is this?
~22% of human proteins have an odd number of cysteines, meaning at least one Cys is unpaired
For E. coli-expressed proteins with native disulfides, incomplete disulfide formation leaves some Cys free
Antibodies have 16+ cysteines; incomplete oxidation creates heterogeneity
When to Mutate Cysteines Out
Mutate to serine (C→S) when:
The Cys is surface-exposed and not part of a disulfide
The protein aggregates via intermolecular disulfides (confirmed by non-reducing SDS-PAGE showing higher MW bands)
The Cys is not functionally important (no catalytic role, no metal coordination)
Mutate to alanine (C→A) when:
The Cys is buried (serine's hydroxyl may be too polar for a hydrophobic environment)
You want to minimize steric changes
Do NOT mutate when:
The Cys coordinates a metal ion (Zn fingers, Fe-S clusters)
The Cys is in the active site (catalytic cysteines)
The Cys forms a structural disulfide that you want to preserve
The Cys is a known PTM site (e.g., palmitoylation, SUMOylation)
Example: Antibody Fragment Engineering
Single-chain variable fragments (scFvs) have 4 cysteines forming 2 intra-domain disulfides (one in VH, one in VL). In E. coli:
Cytoplasmic expression → no disulfides form → aggregation
Periplasmic expression → disulfides form but yield is low
Engineering approach: Express in SHuffle strain OR add a non-natural inter-domain disulfide to stabilize the VH-VL interface → often a better solution than struggling with periplasmic export
When to REDESIGN a Disulfide Bond
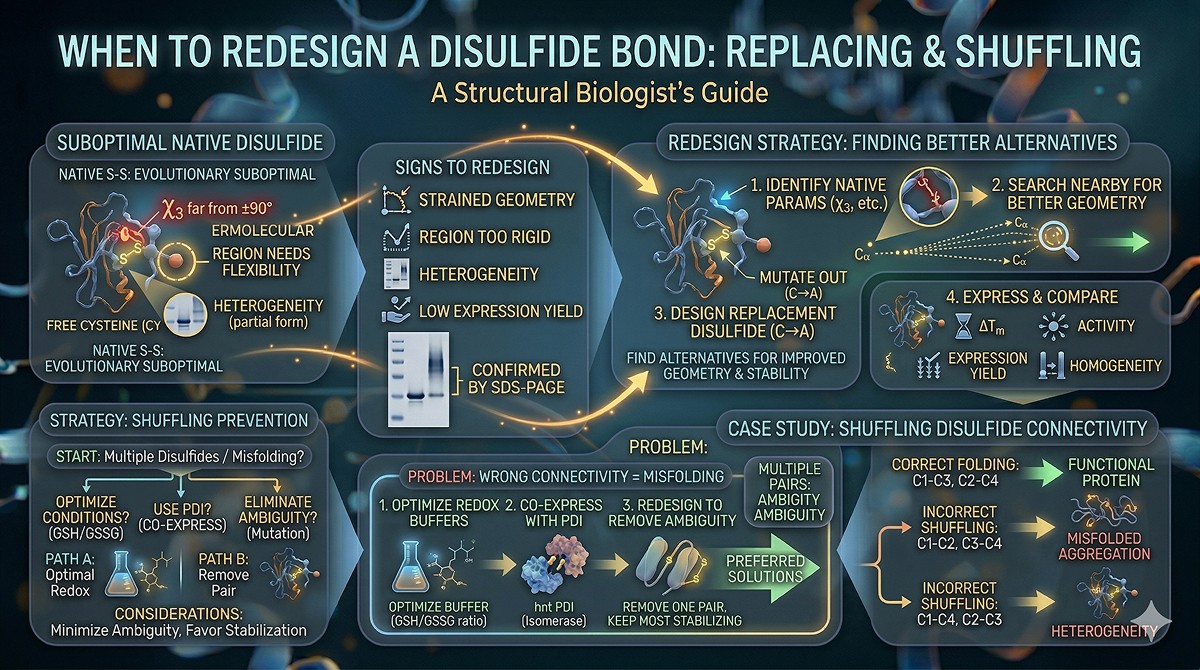
Replacing Non-Ideal Native Disulfides
Some natural disulfides are suboptimal—they formed evolutionarily under different selection pressures than your engineering goals:
Signs a native disulfide should be redesigned:
Strained geometry (χ3 far from ±90°)
Located in a region you need to be flexible (e.g., near an active site hinge)
Creates heterogeneity (partially formed, multiple oxidation states)
Limits expression yield (requires oxidizing conditions that are hard to achieve)
Redesign strategy:
Identify the native disulfide's geometric parameters
Search nearby positions for better-geometry alternatives
Design the replacement disulfide
Express and compare: ΔTm, activity, expression yield, homogeneity
Shuffling Disulfide Connectivity
Some proteins have multiple disulfides that can form in the wrong order:
Correct: C1-C3, C2-C4
Incorrect: C1-C2, C3-C4 (or C1-C4, C2-C3)
Wrong disulfide connectivity = misfolded protein. Solutions:
Optimize oxidative folding conditions (redox buffers: GSH/GSSG ratios)
Co-express with protein disulfide isomerase (PDI)
Redesign to remove ambiguity (if possible, eliminate one pair and keep the most stabilizing one)
Expression System Considerations
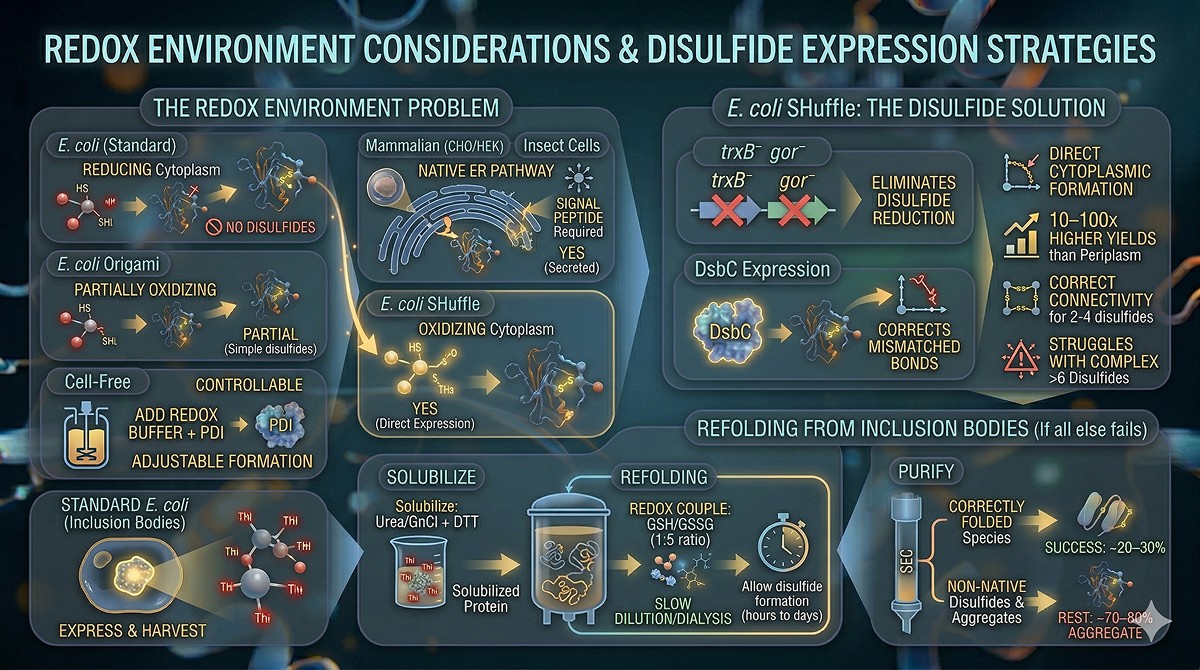
The Redox Environment Problem
Expression System | Cytoplasmic Redox | Disulfide Formation? | Strategy |
|---|---|---|---|
E. coli (standard) | Reducing | No (cytoplasm) | Periplasm, inclusion body refolding |
E. coli SHuffle | Oxidizing cytoplasm | Yes | Direct cytoplasmic expression |
E. coli Origami | Partially oxidizing | Partial | Good for simple disulfides |
Insect cells | Native ER pathway | Yes (secreted) | Signal peptide required |
CHO/HEK293 | Native ER pathway | Yes (secreted) | Signal peptide required |
Cell-free | Controllable | Adjustable | Add redox buffer + PDI |
E. coli SHuffle: The Disulfide Solution
SHuffle strains have two key genetic modifications:
trxB⁻ gor⁻: Eliminates cytoplasmic disulfide reduction (thioredoxin and glutaredoxin pathways knocked out)
DsbC expression: Cytoplasmic expression of disulfide isomerase to correct mismatched disulfides
Results:
Disulfide formation in the cytoplasm
Higher yields than periplasmic expression (10–100x)
Correct disulfide connectivity for proteins with 2–4 disulfides
May struggle with proteins requiring >6 disulfides (complex connectivity)
Refolding from Inclusion Bodies
If all else fails, express in standard E. coli (inclusion bodies) and refold:
Solubilize inclusion bodies in 6–8 M urea or guanidinium chloride + DTT
Slowly dilute or dialyze into refolding buffer
Include a redox couple: oxidized/reduced glutathione (GSSG/GSH, typically 1:5 ratio) or cysteine/cystamine
Allow disulfide formation during refolding (hours to days)
Purify correctly folded species by SEC or ion exchange
Success rate: Approximately 20–30% of proteins with disulfides can be refolded to active material. The rest form non-native disulfides and aggregate.
Computational Tools for Disulfide Engineering
Prediction and Design Tools
Tool | What It Does | Input | Use Case |
|---|---|---|---|
DbD2 (Disulfide by Design) | Identifies residue pairs suitable for engineered disulfides | PDB structure | Design new disulfides |
MODIP | Evaluates structural feasibility of disulfide bonds | PDB structure | Score candidate pairs |
Rosetta DisulfideDesign | Energy-minimized disulfide modeling | PDB structure | Most accurate prediction |
FoldX | ΔΔG of Cys mutations + disulfide bond energy | PDB structure | Quick stability estimate |
AlphaFold2 | Predict mutant structure to check if disulfide distorts fold | Sequence | Validate post-design |
The Workflow
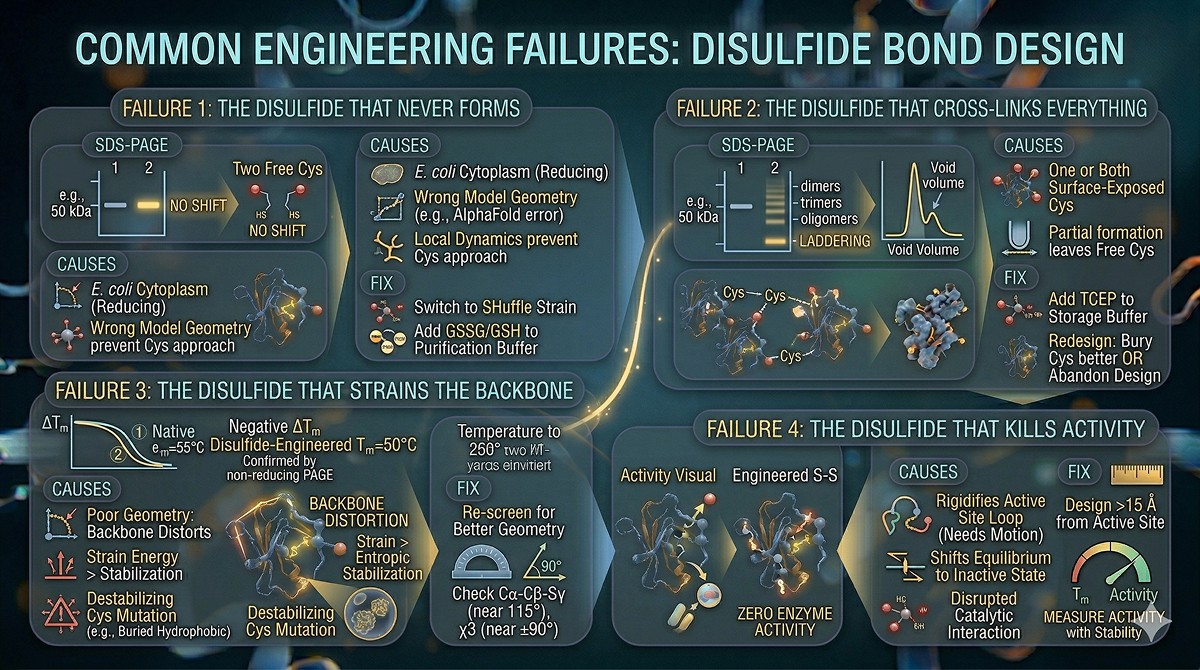
Common Engineering Failures
Failure 1: The Disulfide That Never Forms
Symptom: SDS-PAGE under non-reducing conditions shows no mobility shift compared to reduced. The two cysteines are there but not bonded.
Causes:
Expression in reducing environment (E. coli cytoplasm)
Geometry was wrong in the model (AlphaFold model had error in that region)
Local dynamics prevent the two Cys from approaching each other
Fix: Switch to SHuffle strain, or add GSSG/GSH redox couple to the purification buffer.
Failure 2: The Disulfide That Cross-Links Everything
Symptom: Non-reducing SDS-PAGE shows laddering (dimers, trimers, higher oligomers). SEC shows void volume peak.
Causes:
One or both cysteines are surface-exposed and form intermolecular bonds
Partial disulfide formation leaves free Cys to cross-link with other molecules
Fix: Add TCEP to storage buffers. Or redesign: bury the Cys pair better, or abandon the design.
Failure 3: The Disulfide That Strains the Backbone
Symptom: Disulfide forms (confirmed by non-reducing PAGE) but ΔTm is negative. The protein is less stable with the disulfide.
Causes:
Poor geometry: the backbone must distort to accommodate the disulfide
The strain energy exceeds the entropic stabilization
The Cys mutations themselves are destabilizing (e.g., replacing a buried hydrophobic with polar Cys)
Fix: Re-screen for pairs with better geometry. Ensure Cα-Cβ-Sγ angles are near 115° and χ3 is near ±90°.
Failure 4: The Disulfide That Kills Activity
Symptom: Stable protein, no aggregation—but zero enzyme activity.
Causes:
Disulfide rigidifies an active site loop that needs to move during catalysis
Disulfide shifts the equilibrium between active and inactive conformations
Cys mutation disrupted a catalytic interaction
Fix: Design disulfides >15 Å from the active site. Always measure activity alongside stability.
The Bottom Line
Scenario | Action | Key Consideration |
|---|---|---|
Protein needs more stability, has compatible geometry | Add disulfide | Verify geometry beyond Cβ-Cβ distance; use SHuffle for expression |
Free cysteines cause aggregation | Remove Cys (C→S or C→A) | Verify Cys is not functional |
Native disulfide limits expression | Consider removing it | Test stability and activity without it |
Multiple disulfides form incorrectly | Redesign or eliminate one pair | Simplify disulfide connectivity |
Protein for therapeutic use | Minimize free Cys, optimize connectivity | Homogeneity and stability are both critical |
The golden rule: A well-placed disulfide stabilizes by +5–10°C. A badly placed one destroys the protein. The margin between success and failure is 1–2 Å of geometry and the right expression system.
Disulfide Engineering with Orbion
Orbion's Stabilize module scores any cysteine mutation with AstraDTM (ΔTm) and AstraDDG (ΔΔG), letting you computationally pre-screen disulfide candidates before committing to cloning. AstraPTM predicts native disulfide bonds and other cysteine modifications, flagging residues you shouldn't mutate. AstraUNFOLD provides per-residue disorder and amyloidogenicity tracks—helping you identify regions where disulfide-mediated rigidification might actually help versus regions where flexibility is functional.
For top candidates, AlphaFold2 structure prediction of the variant validates that the engineered disulfide doesn't distort the backbone (via RMSD comparison to wild-type and pocket analysis). Combined with the Construct Design module for choosing the right expression system (SHuffle, periplasmic, or eukaryotic), you can design, score, and plan the expression of disulfide-engineered variants in a single workflow.
References
Dombkowski AA, et al. (2014). Protein disulfide engineering. FEBS Letters, 588(2):206-212. Link
Pace CN, et al. (1988). Contribution of the hydrophobic effect to globular protein stability. Journal of Biological Chemistry, 263(24):11820-11825. Link
Lobstein J, et al. (2012). SHuffle, a novel Escherichia coli protein expression strain capable of correctly folding disulfide bonded proteins in its cytoplasm. Microbial Cell Factories, 11:56. Link
Matsumura M, et al. (1989). Substantial increase of protein stability by multiple disulphide bonds. Nature, 342:291-293. Link
Fass D. (2012). Disulfide bonding in protein biophysics. Annual Review of Biophysics, 41:63-79. Link
Craig DB & Bhatt S. (2008). Disulfide by Design 2.0: a web-based tool for disulfide engineering in proteins. BMC Bioinformatics, 14:346. Link
Singh A, et al. (2015). Protein recovery from inclusion bodies of Escherichia coli using mild solubilization process. Microbial Cell Factories, 14:41. PMC4030991
Trivedi MV, et al. (2009). The role of thiols and disulfides in protein chemical and physical stability. Current Protein and Peptide Science, 10(6):614-625. PMC2802230
Wong JWH, et al. (2011). Disulfide bond acquisition through eukaryotic protein evolution. Molecular Biology and Evolution, 28(1):327-334. Link
Berkmen M. (2012). Production of disulfide-bonded proteins in Escherichia coli. Protein Expression and Purification, 82(1):240-251. Link
