Blog
Orbion Team
My AlphaFold Model Doesn't Match My Crystal Structure

You solved your crystal structure at 2.0 Å. You also ran AlphaFold2. The backbone RMSD between them is 3.5 Å. The loops are in different places. A helix is shifted. One domain seems rotated. Which one is right—the experimental structure or the computational prediction?
Usually, both are right. They're just showing you different things. Understanding why AlphaFold and crystal structures disagree is essential for anyone using structural models to guide experiments.
Key Takeaways
AlphaFold predicts a single static model—it doesn't capture the conformational ensemble your protein actually samples
Crystal structures are biased by crystal packing—contacts between symmetry mates can distort loops, shift domains, and trap rare conformations
Most disagreements are in loops and flexible regions—check the pLDDT and PAE before assuming AlphaFold is wrong
Domain orientations are a known weakness—AlphaFold may predict individual domains well but get the relative orientation wrong (check PAE inter-domain)
Neither model is "ground truth"—the truth is a conformational ensemble, and both models are snapshots
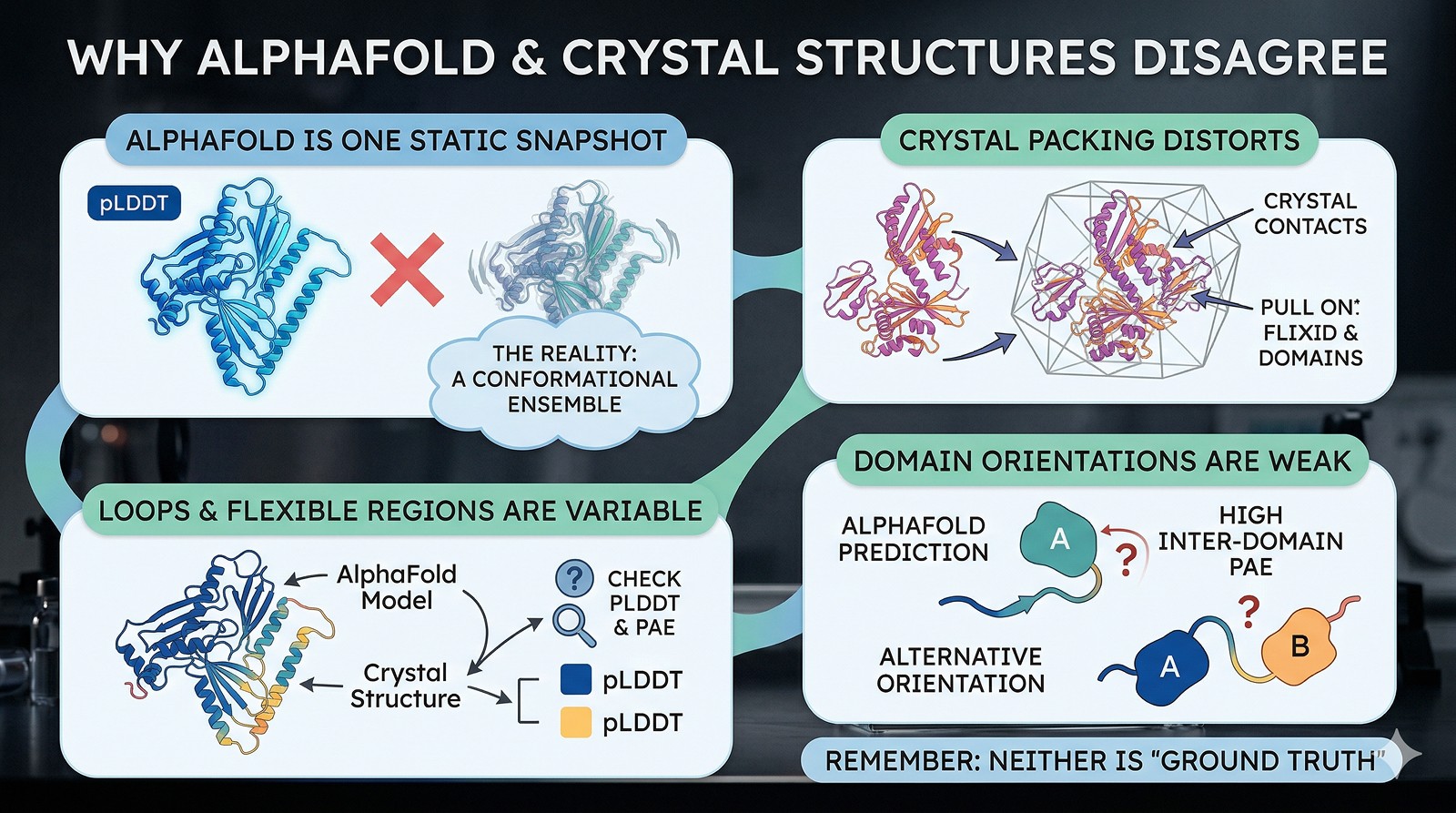
Why They Disagree
Reason 1: Crystal Packing Artifacts
Crystal structures are solved in a crystalline lattice. Protein molecules make contacts with symmetry-related neighbors that don't exist in solution. These contacts can:
Pin flexible loops in a single conformation (that may be rare in solution)
Shift domain orientations by 5–15° compared to the solution state
Select rare conformations that happen to pack well in the lattice
How to check: Use PDBe PISA or similar tools to identify crystal contacts. If the disagreement region is involved in a crystal contact, the crystal structure is biased—not AlphaFold.
Reason 2: Flexible Regions and Conformational Ensembles
AlphaFold gives you one model. Your protein in solution samples many conformations. The crystal structure captures one conformation (the one that crystallized). These two single conformations may differ—and both may be valid members of the ensemble.
AlphaFold's signal: pLDDT < 70 in a region means AlphaFold is uncertain. The predicted structure there shouldn't be trusted
B-factors in the crystal structure: High B-factors (>60 Ų) mean that region is flexible even in the crystal. The coordinates are imprecise
The rule: If both AlphaFold (low pLDDT) and the crystal structure (high B-factors) flag a region as uncertain, that region is genuinely flexible—neither model is capturing the full picture.
Reason 3: Ligand-Induced Conformational Changes
AlphaFold predicts the apo state (no ligand). Your crystal structure may have been solved with a ligand, substrate, inhibitor, or cofactor bound. Ligand binding commonly causes:
Loop ordering (disordered → ordered upon binding)
Domain closure (open → closed upon substrate binding)
Allosteric rearrangements
How to check: Was there anything in the crystallization condition? Co-crystallization with a ligand? Even buffer components (citrate, sulfate, PEG fragments) can bind to active sites and shift conformations.
Reason 4: Multi-Domain Relative Orientation
AlphaFold predicts individual domains well (backbone RMSD < 1 Å for most globular domains). But the relative orientation between domains connected by flexible linkers is a known weakness.
How to check: Align each domain separately. If individual domain RMSDs are < 1.5 Å but the full-chain RMSD is > 3 Å, the problem is inter-domain orientation, not fold prediction
PAE tells you this: If the PAE matrix shows low confidence (high error) between domains, AlphaFold is explicitly telling you it doesn't know the relative orientation
Reason 5: Oligomeric State Effects
AlphaFold2 predicts monomers by default. If your crystal structure is a dimer or higher oligomer, subunit interfaces can shift domain positions and stabilize conformations that don't exist in the monomer.
Fix: Run AlphaFold-Multimer with the correct stoichiometry. The predicted complex may match the crystal structure much better
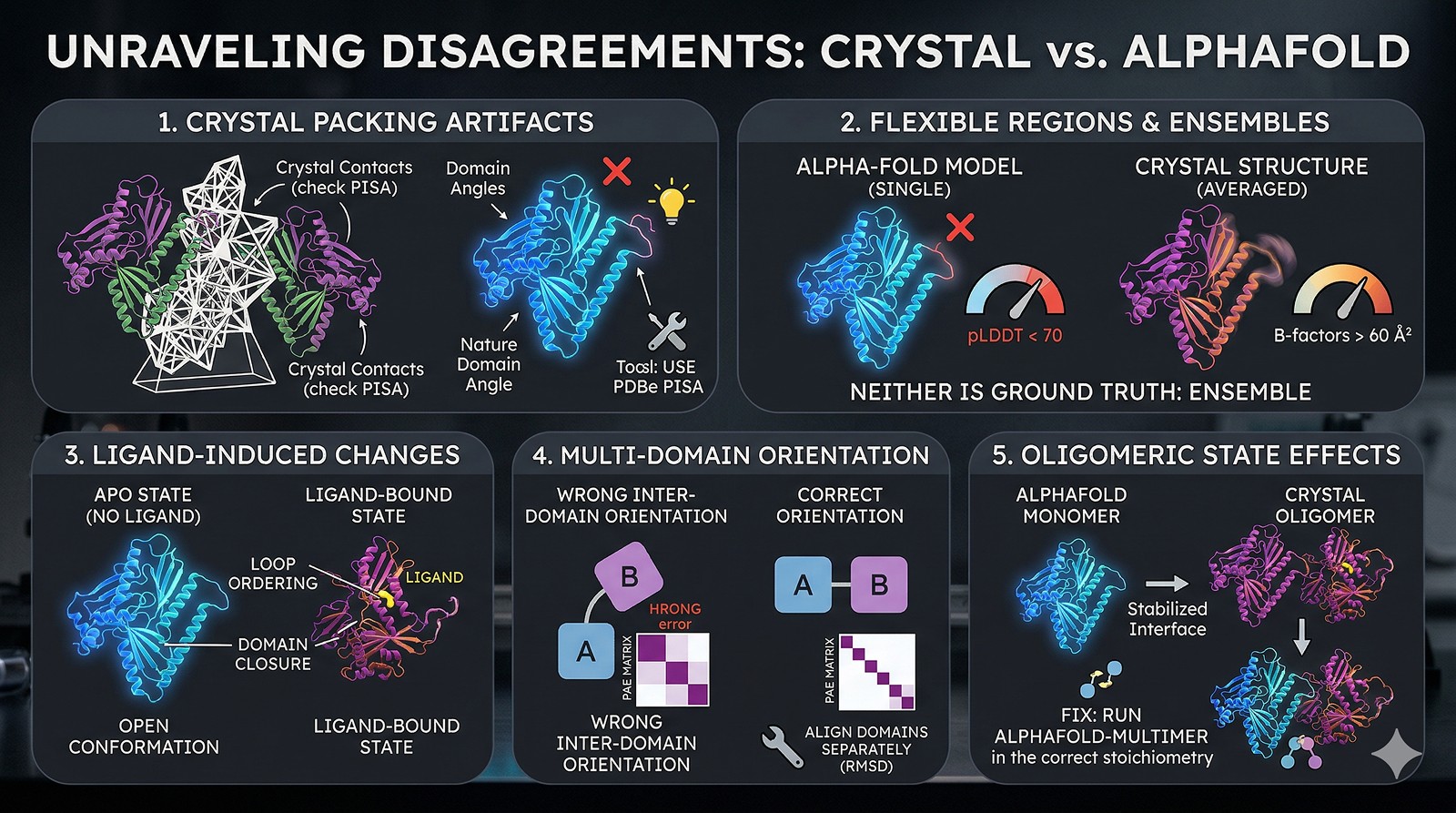

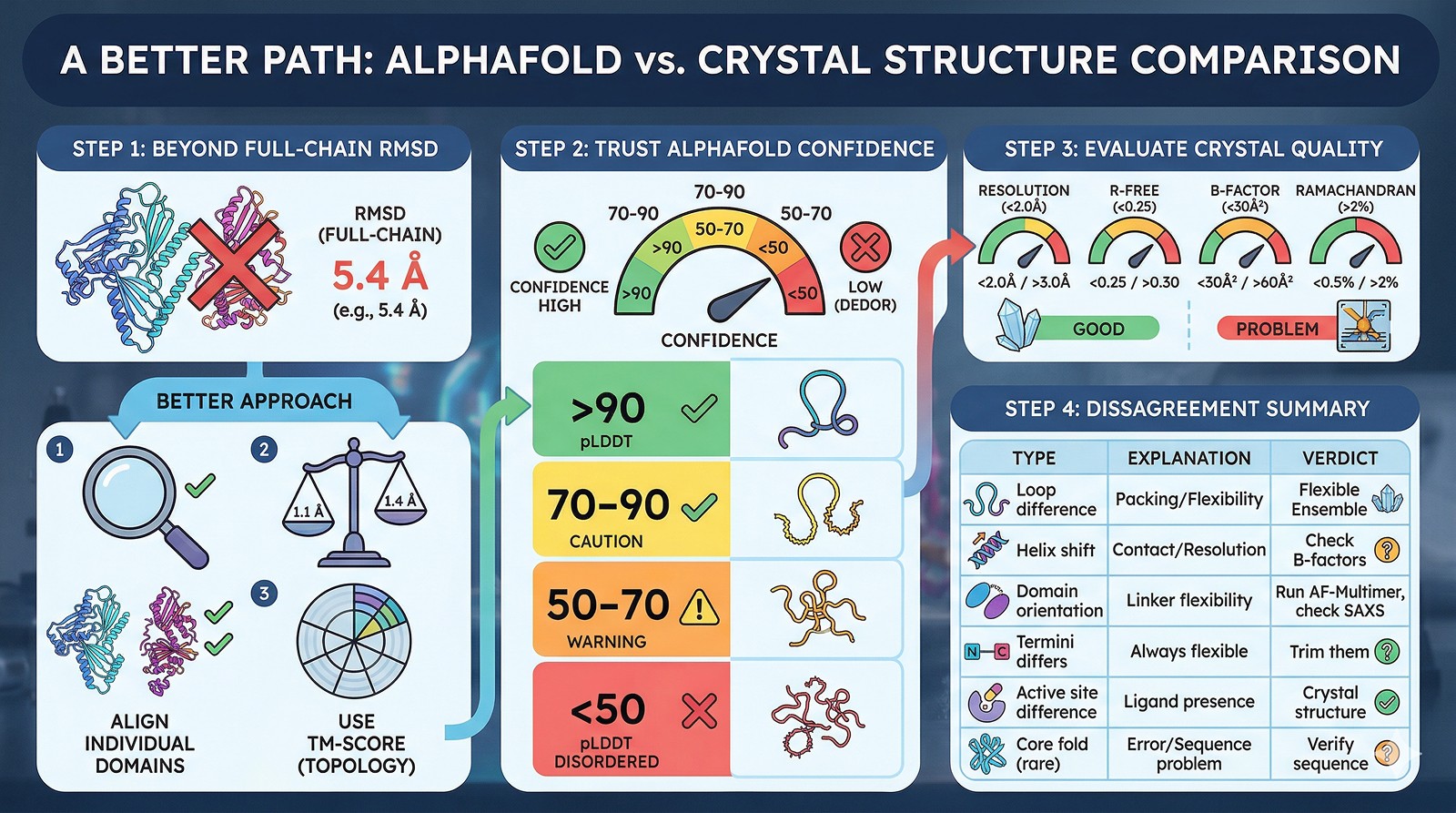
How to Compare Properly
Step 1: Don't Use Full-Chain RMSD
Full-chain RMSD is a terrible metric for multi-domain proteins or proteins with flexible termini. It's dominated by the worst-aligned region.
Better approach:
Align individual domains (or secondary structure elements) separately
Report per-domain RMSD
Use TM-score (topology-based) instead of RMSD—it's less sensitive to local outliers
Step 2: Check AlphaFold Confidence
pLDDT Range | Interpretation | Trust Level |
|---|---|---|
>90 | Very high confidence | AlphaFold likely correct |
70–90 | Confident | Generally reliable; minor deviations expected |
50–70 | Low confidence | Don't trust coordinates; region is likely flexible |
<50 | Very low / disordered | Likely intrinsically disordered; no defined structure |
Step 3: Check Crystal Structure Quality
Metric | Good | Caution | Problem |
|---|---|---|---|
Resolution | <2.0 Å | 2.0–3.0 Å | >3.0 Å |
R-free | <0.25 | 0.25–0.30 | >0.30 |
B-factor (region) | <30 Ų | 30–60 Ų | >60 Ų |
Ramachandran outliers | <0.5% | 0.5–2% | >2% |
Step 4: The Comparison Table
Disagreement Type | Likely Explanation | Who's Probably Right |
|---|---|---|
Loop conformation differs | Crystal packing or genuine flexibility | Neither—it's flexible |
Helix shifted by 1–2 Å | Crystal contact or resolution limit | Check B-factors and packing |
Domain orientation differs | Flexible linker; different conformational state | Run AF-Multimer; check SAXS data |
N/C-terminus differs | These are always flexible | Neither—trim them for comparison |
Active site differs | Ligand in crystal structure | Crystal structure (if ligand present) |
Core fold differs (rare) | Possible AlphaFold error or wrong sequence | Verify sequence; check MSA depth |
When AlphaFold Is Actually Wrong
AlphaFold does make genuine errors. Watch for:
Shallow MSAs (few homologs in the alignment): prediction quality drops significantly
Novel folds not well represented in the training set
Transmembrane proteins in detergent micelles vs lipid bilayer conformations
Metalloproteins where metal coordination organizes the structure (AlphaFold doesn't model metals)
The tell: If pLDDT is high (>85) but the structure still doesn't match, investigate carefully. High-confidence AlphaFold errors are rare but do occur, especially for proteins with few homologs.
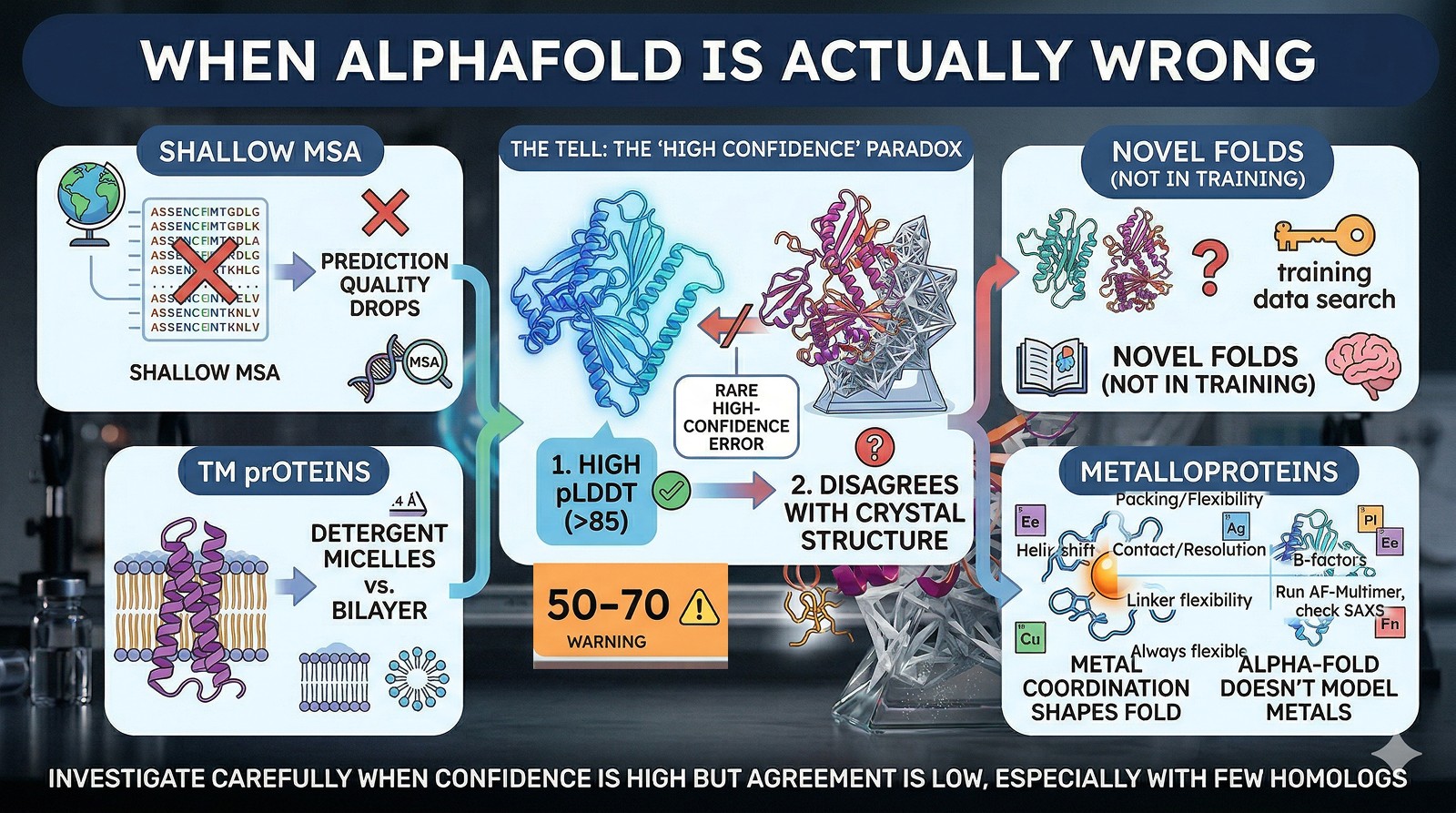

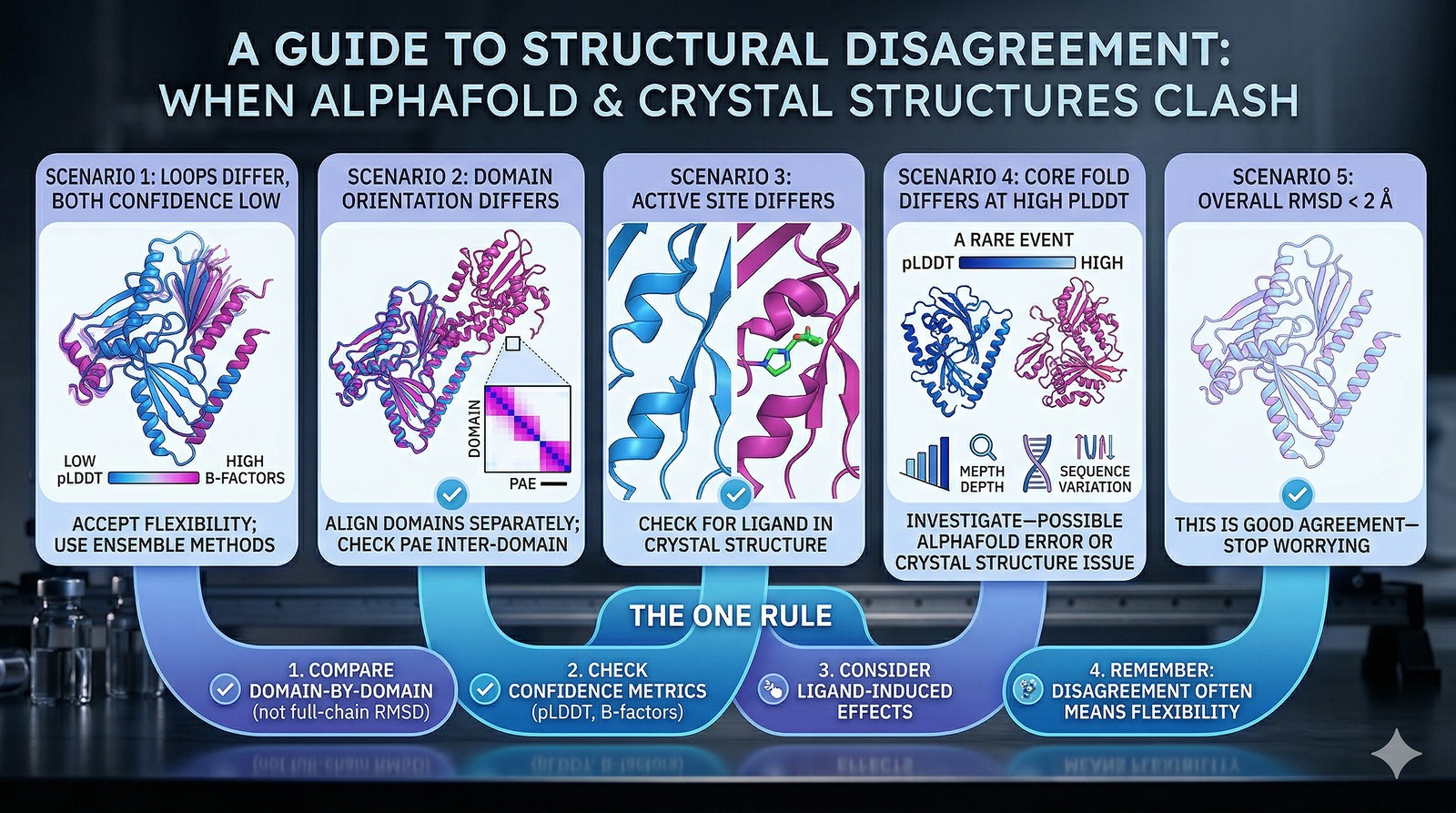
The Bottom Line
Scenario | What to Do |
|---|---|
Loops differ, both have low confidence/high B-factors | Accept flexibility; use ensemble methods |
Domain orientation differs | Align domains separately; check PAE inter-domain |
Active site conformation differs | Check for ligand in crystal structure |
Core fold differs at high pLDDT | Investigate—possible AlphaFold error or crystal structure issue |
Overall RMSD < 2 Å | This is good agreement—stop worrying |
The one rule: Never compare a full-chain RMSD and declare one model "wrong." Compare domain by domain, check confidence metrics on both sides, and remember that disagreement often means flexibility—not error.
Using Orbion for Structural Comparison
Orbion integrates AlphaFold2 structure prediction with PAE Insight Engine analysis, making it straightforward to identify which regions of your model are high-confidence and which are uncertain. The PAE inter-domain analysis specifically highlights where domain orientations should not be trusted—exactly the regions where AlphaFold and crystal structures most commonly disagree. Combined with AstraUNFOLD's disorder predictions, you can distinguish genuine structural differences from regions that are simply too flexible for any single model to capture.
References
Jumper J, et al. (2021). Highly accurate protein structure prediction with AlphaFold. Nature, 596:583-589. Link
Thornton JM, et al. (2021). AlphaFold heralds a data-driven revolution in biology and medicine. Nature Medicine, 27:539-540. Link
Zhang Y, Skolnick J. (2004). Scoring function for automated assessment of protein structure template quality. Proteins, 57(4):702-710. Link
