Blog
Orbion Team
SDS-PAGE Band at the Wrong Molecular Weight

Your protein is 45 kDa. You calculated it from the sequence. The mass spec confirms it. But on SDS-PAGE, it runs at 55 kDa. Or 35 kDa. Or as a doublet. The ladder doesn't lie—but neither does mass spectrometry. So what's going on?
SDS-PAGE is the most widely used protein analysis technique in biology, and it's wrong more often than most people realize. Not broken—just measuring something different from what you think.
Key Takeaways
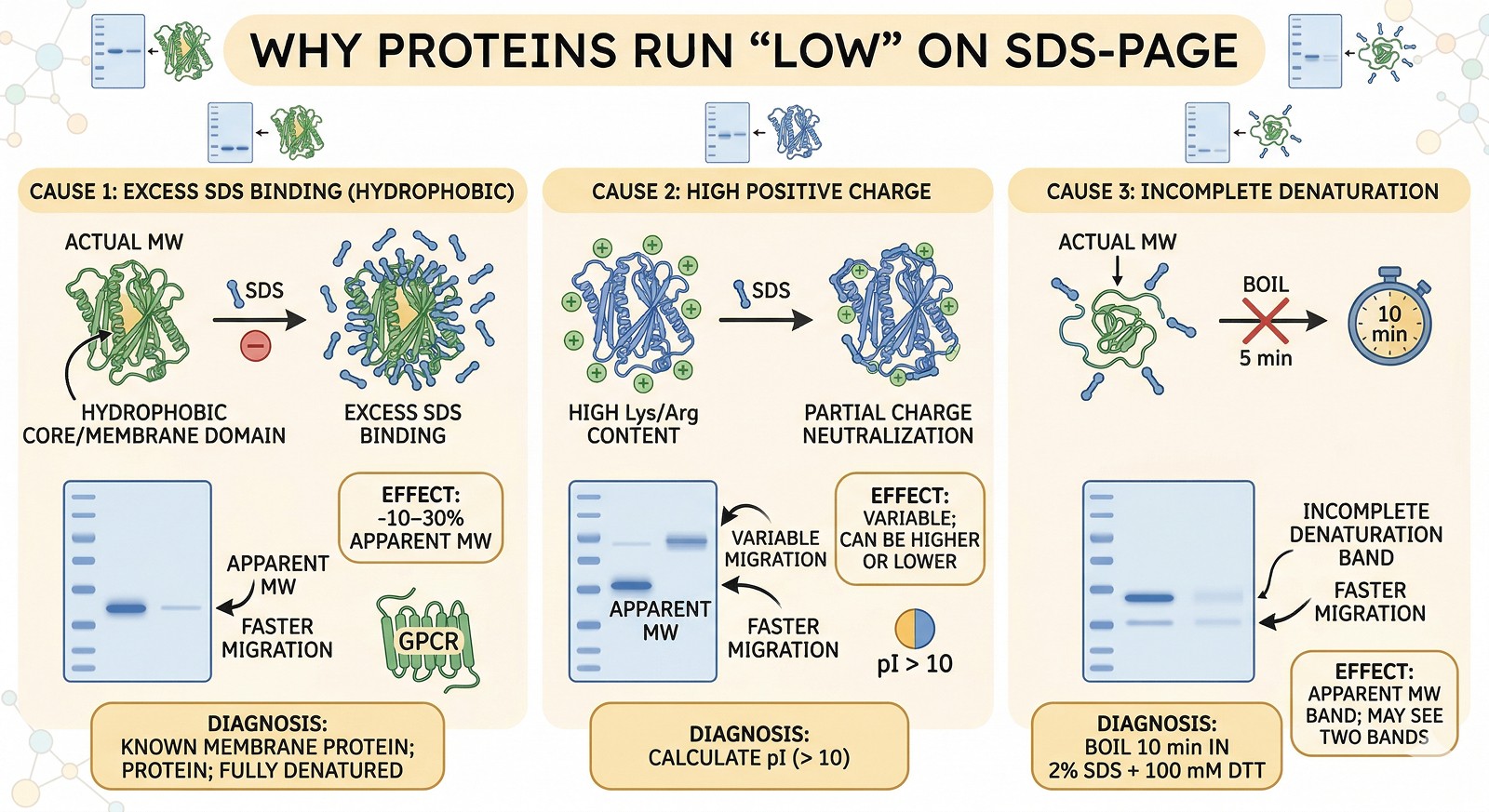
SDS-PAGE measures electrophoretic mobility, not molecular weight—anomalous migration is common and usually explainable
Running high: glycosylation, high negative charge, SDS-resistant structure, proline-rich sequences, and intrinsic disorder all cause slow migration
Running low: highly positively charged proteins, hydrophobic proteins that bind excess SDS, and membrane proteins with bound lipid all migrate faster
Doublets usually mean partial modification: incomplete PTMs, partial proteolysis, or partial reduction of disulfides
Mass spectrometry is the ground truth for molecular weight—SDS-PAGE is an estimate at best
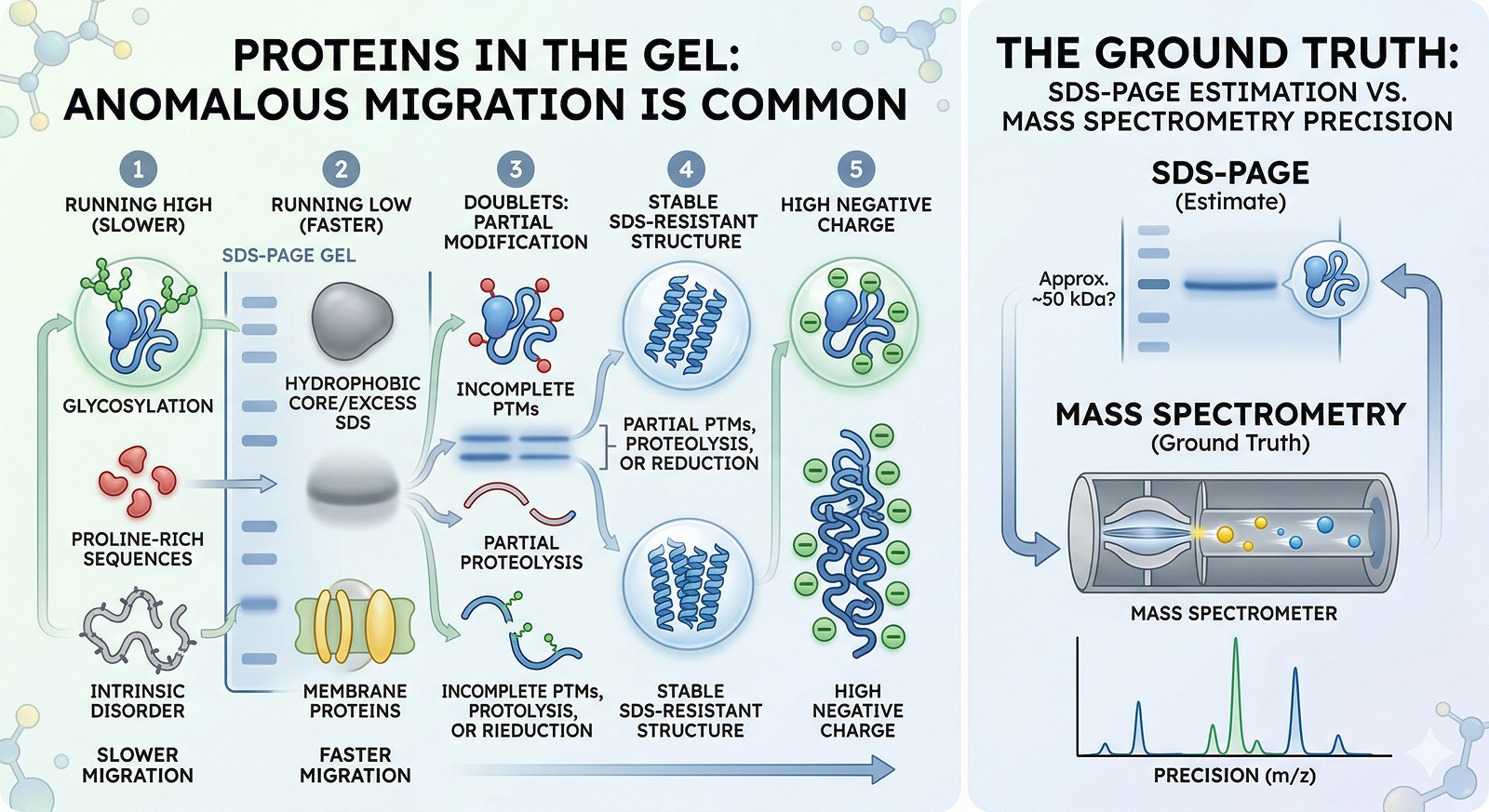
Why SDS-PAGE Gets the MW "Wrong"
How SDS-PAGE Works
SDS (sodium dodecyl sulfate) binds to proteins at a roughly constant ratio (~1.4 g SDS per gram of protein), unfolding them into linear rods with uniform negative charge density. In theory, all proteins then migrate purely by size through the polyacrylamide gel.
In practice, this works for ~70% of proteins. For the other 30%, the assumptions break down.
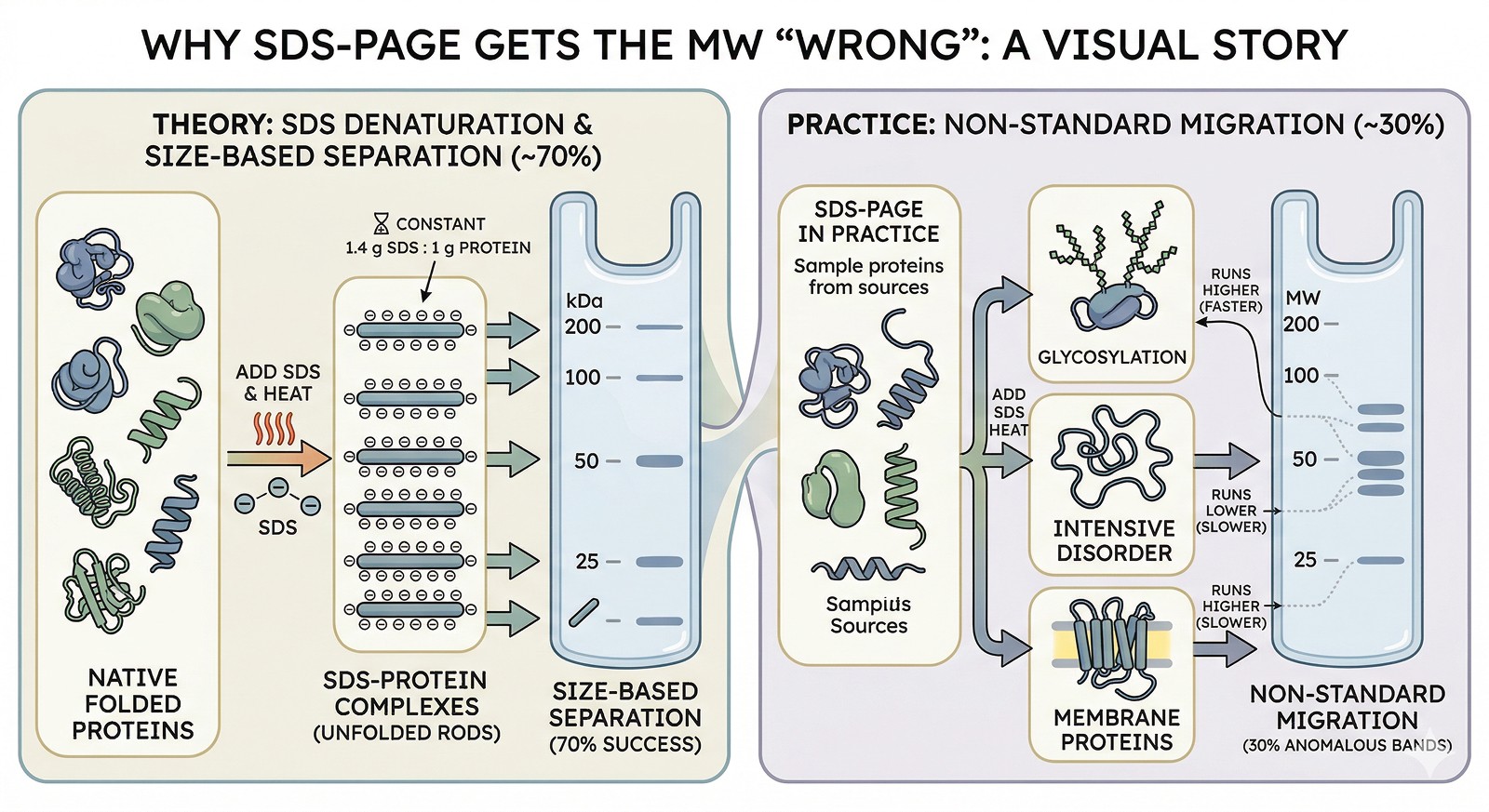
Running HIGH (Apparent MW > Actual MW)
Cause 1: Glycosylation
Glycans don't bind SDS. Glycosylated proteins have reduced charge-to-mass ratio → slower migration → appear heavier.
Effect: +5–30 kDa apparent increase depending on glycan content
Diagnosis: Treat with PNGase F (removes N-glycans); band shifts down to expected MW
Classic example: Glycoproteins often appear as broad, diffuse bands ("smears") due to glycan heterogeneity
Cause 2: Intrinsic Disorder / Proline-Rich Sequences
Disordered proteins and proline-rich regions don't fully linearize in SDS. The extended conformation creates a larger effective size.
Effect: +10–50% apparent MW increase
Diagnosis: AlphaFold pLDDT < 50 for large portions; mass spec confirms true MW
Classic example: Tau protein (45 kDa) runs at ~60–70 kDa on SDS-PAGE
Cause 3: High Negative Charge (Asp/Glu-Rich)
Proteins with many acidic residues already carry more negative charge than average. SDS adds less incremental charge → lower charge-to-mass ratio → slower migration.
Effect: +5–15% apparent MW increase
Diagnosis: Calculate net charge at pH 7; high Asp+Glu content (>15%)
Cause 4: SDS-Resistant Structure
Some protein domains resist SDS denaturation, maintaining a compact structure that migrates slower than the linear form.
Effect: Variable (can be dramatic)
Diagnosis: Boil sample for 10 min (vs standard 5 min at 95°C); or try 8 M urea + SDS
Classic example: Outer membrane proteins (OMPs) often run at a different MW when boiled vs unboiled
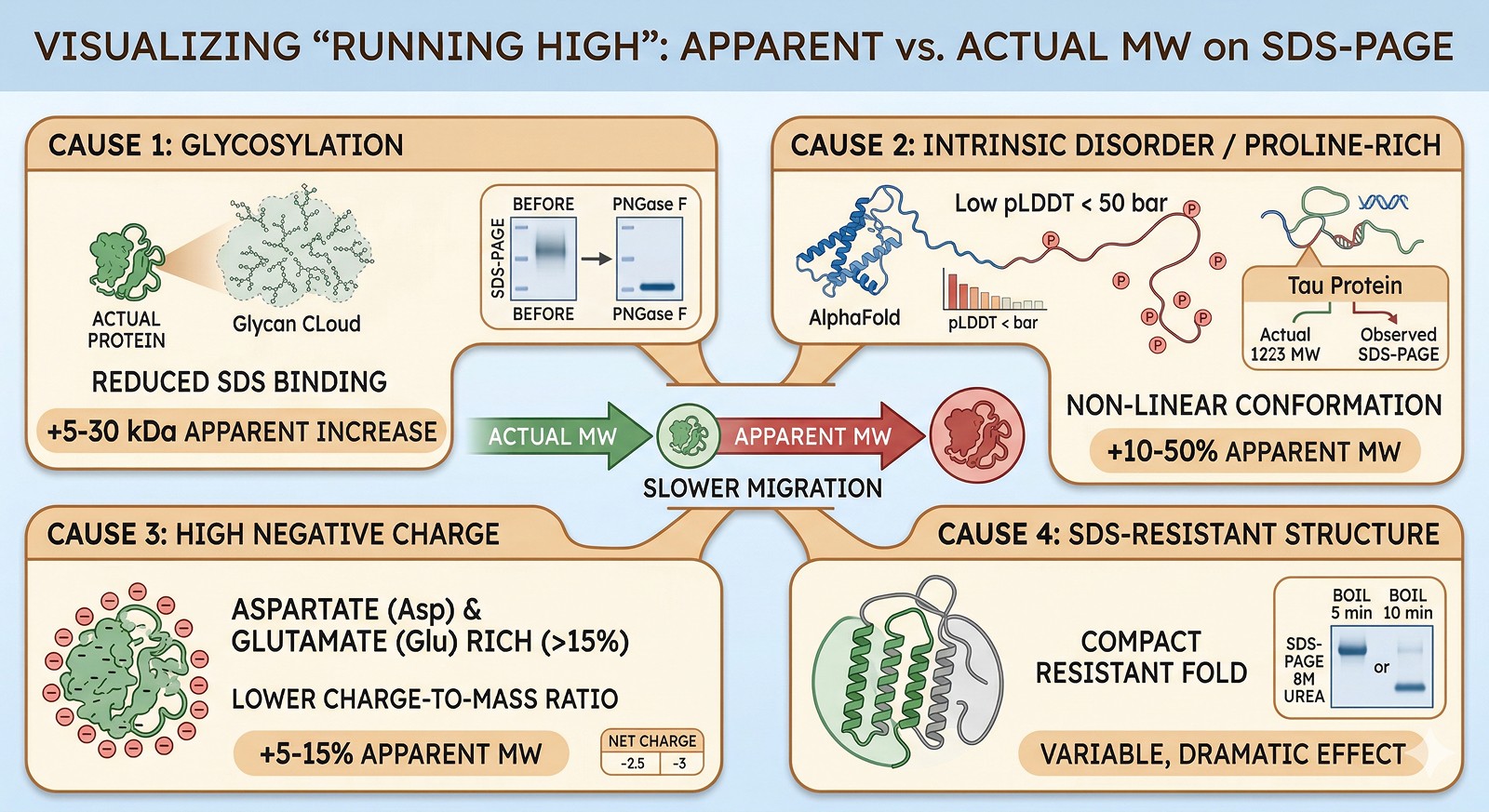
Running LOW (Apparent MW < Actual MW)
Cause 1: Excess SDS Binding (Hydrophobic Proteins)
Highly hydrophobic proteins (especially membrane proteins) bind more SDS than average → higher charge-to-mass ratio → faster migration.
Effect: –10–30% apparent MW
Diagnosis: Known membrane protein; runs lower than expected even when fully denatured
Classic example: Many GPCRs and transporters run 20–30% below predicted MW
Cause 2: Highly Positively Charged Proteins
Very basic proteins (high Lys/Arg content) partially neutralize the SDS negative charge → unusual migration.
Effect: Variable; can run higher or lower depending on the specific protein
Diagnosis: Calculate pI; very basic proteins (pI > 10) often misbehave on SDS-PAGE
Cause 3: Incomplete Denaturation
If the protein isn't fully denatured, it retains some compact structure → faster migration than the fully linearized form.
Effect: Band at lower apparent MW; may see two bands (native + denatured)
Fix: Ensure full denaturation: boil 10 min in 2% SDS + 100 mM DTT
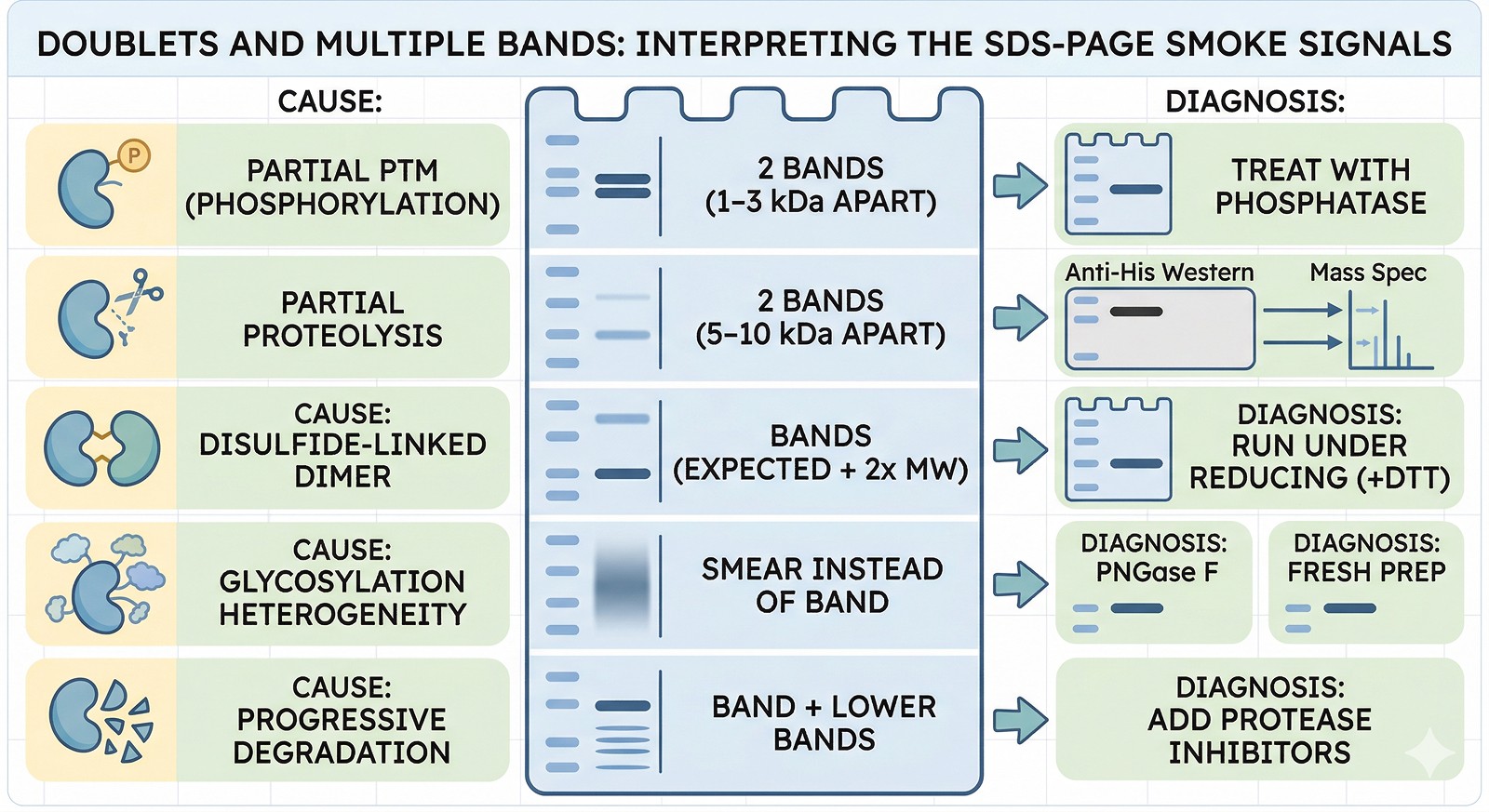
Doublets and Multiple Bands
Pattern | Most Likely Cause | Diagnosis |
|---|---|---|
Two bands, 1–3 kDa apart | Partial PTM (phosphorylation, glycosylation) | Treat with phosphatase or PNGase F; one band disappears |
Two bands, 5–10 kDa apart | Partial proteolysis | Anti-His Western: is the tag on both bands? Mass spec both bands |
Two bands, one at expected MW, one at 2x | Disulfide-linked dimer | Run under reducing (+DTT) conditions: dimer band disappears |
Smear instead of band | Glycosylation heterogeneity or aggregation | PNGase F for glycosylation; fresh prep for aggregation |
Band + multiple lower bands | Progressive degradation | Add protease inhibitors; process faster |
The Bottom Line
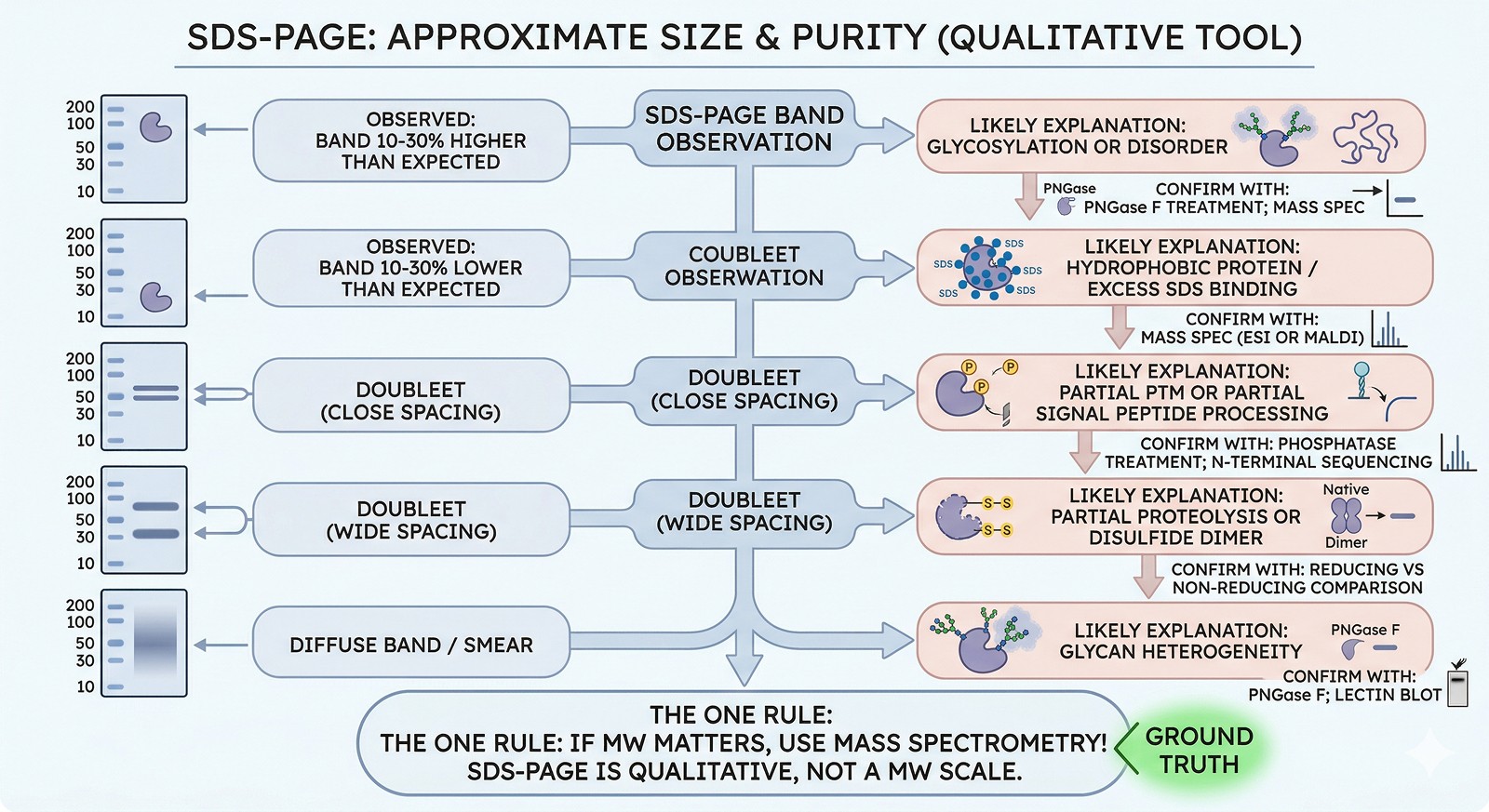
SDS-PAGE Observation | Likely Explanation | Confirm With |
|---|---|---|
Band 10–30% higher than expected | Glycosylation or disorder | PNGase F treatment; mass spec |
Band 10–30% lower than expected | Hydrophobic protein / excess SDS binding | Mass spec (ESI or MALDI) |
Doublet (close spacing) | Partial PTM or partial signal peptide processing | Phosphatase treatment; N-terminal sequencing |
Doublet (wide spacing) | Partial proteolysis or disulfide dimer | Reducing vs non-reducing comparison |
Diffuse band / smear | Glycan heterogeneity | PNGase F; lectin blot |
The one rule: If the MW matters for your experiment, use mass spectrometry. SDS-PAGE tells you approximate size and purity—it's a qualitative tool, not a molecular weight scale.
Predicting the Anomalies
Orbion's AstraPTM predicts glycosylation, phosphorylation, and other modifications that cause anomalous SDS-PAGE migration. AstraUNFOLD identifies disordered regions that run high. Knowing these properties before you run the gel helps you interpret what you see—and avoid chasing phantom "contaminants" that are actually your protein running at the wrong position.
References
Rath A, Glibowicka M, Nadeau VG, Chen G, Deber CM. (2009). Detergent binding explains anomalous SDS-PAGE migration of membrane proteins. PNAS, 106(6):1760-1765. PMC2644116
Tompa P. (2002). Intrinsically unstructured proteins. Trends in Biochemical Sciences, 27(10):527-533. Link
Freeze HH, Kranz C. (2010). Endoglycosidase and glycoamidase release of N-linked glycans. Current Protocols in Molecular Biology, Chapter 17:Unit 17.13A. Link
