Publication
Çağlar Bozkurt
Why Some Binder Signals Transfer and Others Flip

When people evaluate de novo protein binders, the focus usually lands on two things:
Does the binder look structurally plausible?
Does it seem likely to bind the target well?
Those questions matter. But they are not enough.
A binder can look promising in-silico and still fail experimentally. It may not express well. It may aggregate. It may behave differently when deployed in a CAR construct versus as a standalone protein. It may pass one gate and fail another. That is the gap we explored in our new preprint.
In the paper, we re-analysed two public binder-design benchmarks using biology-informed sequence descriptors derived from Orbion’s Astra models. The goal was not to replace structural evaluation, but to test whether there are additional sequence-level signals associated with success or failure that structure- and affinity-centered evaluation does not fully capture.
The Two Datasets We Analysed
We looked at two public benchmarks with very different design contexts.
The first was the Bits to Binders CAR-T CD20 benchmark. In that setting, the binder is used as part of a membrane-displayed CAR construct in primary human T cells. That means the sequence is not just being asked to bind. It also has to function in a multi-part receptor context and survive several experimental gates, including recovery, enrichment, and depletion.
The second was the Adaptyv EGFR benchmark. There, binders are tested as standalone proteins using cell-free expression and bio-layer interferometry. This is a different problem. The binder needs to fold independently and engage a large extracellular target surface without the same architectural constraints as a CAR format.
That difference in deployment context ended up being central.

The Short Version of the Result
We found that the signals associated with binder success fall into three buckets:
Transferable signals
Architecture-dependent signals
Context-specific signals
This matters because it suggests there is no single universal screening logic that works equally well across all binder problems.
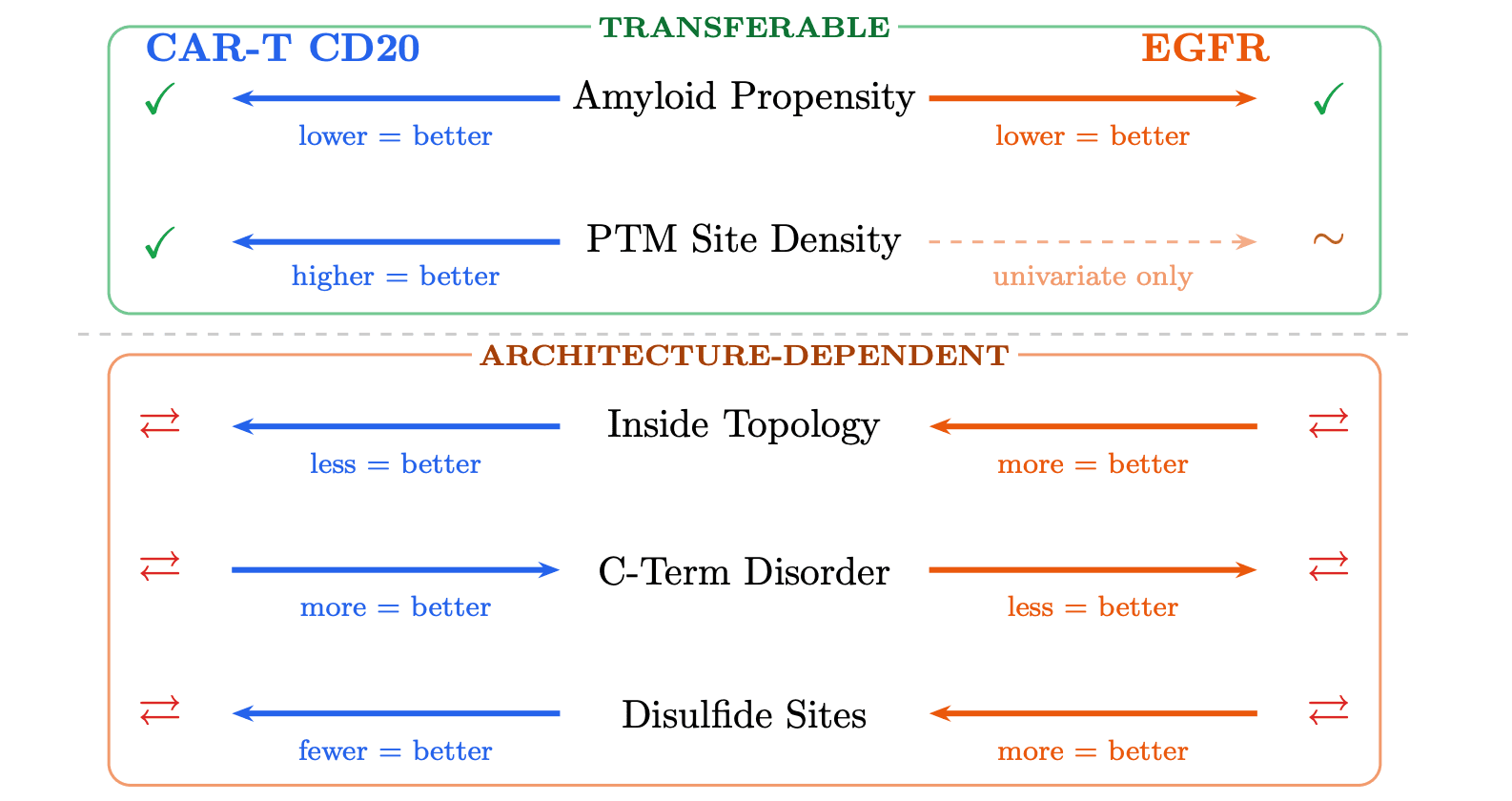
1. Transferable Signals
Some signals showed the same directional association with success across both benchmarks.
The clearest example was lower aggregation propensity. Across both the CAR-T and EGFR datasets, lower amyloid / aggregation-related signal was associated with better outcomes. In other words, this appears to be one of the more robust things worth screening for before synthesis and testing.
We also saw a recurring association with predicted PTM-site density. In univariate analysis, higher PTM-site counts were associated with success in both benchmarks. But that signal was more complicated in EGFR because the dataset includes variable-length binders, so sequence length partly confounds count-based features.
So the cleanest transferable signal in the paper is not “higher affinity-like score.” It is closer to: avoid sequences that look more aggregation-prone.
2. Architecture-dependent Signals
This is where the paper gets more interesting.
Some feature families were significant in both datasets, but flipped direction depending on context. That means the same signal could help in one binder setting and hurt in another.
The strongest examples were:
topology-like character
disorder
disulfide-related features
Why might that happen?
Because a CAR-T binder and a standalone EGFR binder are solving different biological problems.
In the CAR-T benchmark, the binder is part of a membrane-displayed receptor system. Some degree of flexibility or extracellular compatibility may be helpful. In the EGFR benchmark, the binder is a standalone folded unit. There, lower disorder and more compact fold-related character were much more favorable.
The same applies to disulfide-related features. In one context they may look like structural support; in another they may become a misfolding liability.
This is one of the core takeaways of the paper: you should not assume that a useful screening signal in one binder architecture will transfer cleanly into another.
3. Context-specific Signals
Some associations only appeared in one benchmark, or only at one experimental gate.
In the CAR-T dataset, one of the notable findings was a phosphorylation-related depletion association. We also saw an expression versus enrichment tradeoff, where some features flipped direction across gates within the same benchmark. A sequence can look better for one stage of the pipeline and worse for another.
In the EGFR dataset, the strongest signal was low disorder. This stood out more strongly there than any single feature did in the CAR-T analysis.
That is important because it changes how screening should be framed.
Instead of asking only, “Is this a good binder candidate?”, it can be more useful to ask: What kind of failure risk does this sequence carry in this deployment context?
A Practical Result
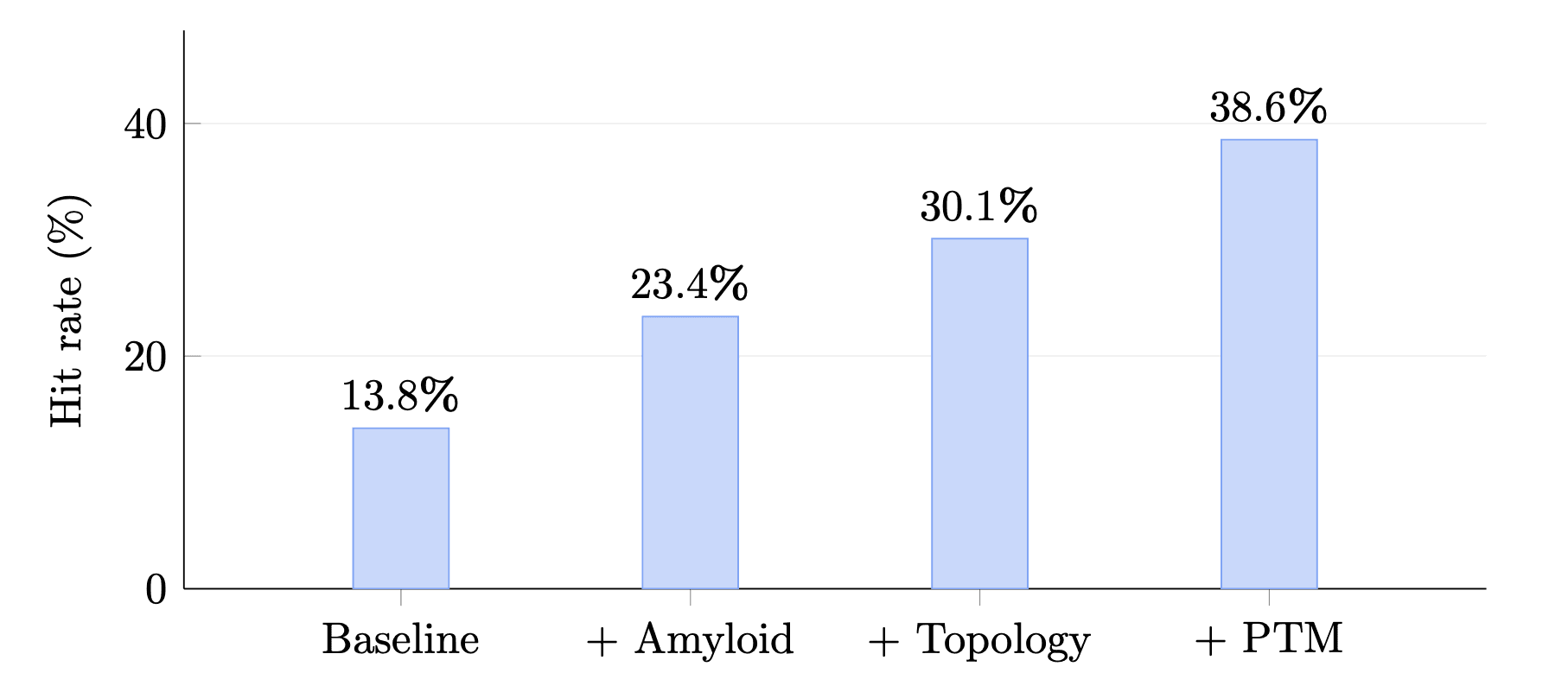
We also tested a simple retrospective filter stack in the CAR-T benchmark.
Starting from a controlled subset, adding biology-informed filters increased the enrichment hit rate from 13.8% to 38.6%, which is a 2.8× lift. The exact stack was specific to that benchmark and was selected retrospectively, so it should not be treated as a production-ready rule set. But it shows that these features can have practical screening value when layered on top of existing heuristics.
This is an important nuance.
The paper does not claim that one fixed filter recipe solves binder evaluation. It argues for something more careful: a layered, context-aware screening approach.
The Broader Idea
The bigger message of the paper is simple: binder success depends on deployment context.
A sequence is not being evaluated in a vacuum. It is being asked to function in a specific architecture, assay format, and biological setting. That means the same descriptor can carry different implications depending on where and how the binder is used.
We think this is a useful shift in perspective.
Instead of treating binder evaluation as a universal ranking problem, it may be better to treat it as a multi-gate, context-aware screening problem. That framing is closer to how experimental failure actually happens.
Important Limitations
This study is a retrospective re-analysis of public benchmark data. It is correlational. We did not synthesize or test new binders as part of this paper. Some signals are stronger than others, and some findings remain tentative, especially where sample sizes were smaller or feature counts were partly length-confounded.
So this is not the final word.
But it does point toward a more useful evaluation layer: one that tries to identify not just whether a design looks strong, but what kind of experimental risk profile it may carry before it reaches the lab.
You can read the full preprint here: https://www.biorxiv.org/content/10.64898/2026.04.13.718094v1
Book a 20-Minute Demo
Sign up free for unlimited Overview runs — summary, sequence-based analysis, homology search. For the full Characterization — PTMs, binding sites, stability variants, construct design — book a demo and we'll run your target live.
