Publication
Çağlar Bozkurt
Beyond Structure and Affinity: Our New Preprint on CD20 and EGFR Benchmarks

De novo protein binder design has advanced quickly.
Generative models can now produce thousands of candidate binders for a target. Structural scoring has improved. Affinity-centered evaluation has become standard. And yet experimental hit rates remain far lower than most teams want.
That gap is what motivated our latest preprint.
Today, we are sharing a new study from Orbion: “Beyond Structure and Affinity: Context-Dependent Signals for de novo Binder Success.” In this work, we re-analysed two public binder benchmarks to ask a simple question:
Are there sequence-level biological signals associated with binder success that structure- and affinity-centered evaluation does not fully capture?
Our answer is: yes — but those signals are not universal. They depend on context.
Using biology-informed sequence descriptors derived from Orbion’s Astra models, we analysed two very different public datasets:
the Bits to Binders CAR-T CD20 benchmark, where binders function as part of a membrane-displayed CAR construct in human T cells
the Adaptyv EGFR benchmark, where binders are tested as standalone proteins in a cell-free expression and binding setup
These are not the same design problem. And that difference turns out to matter.
Across the two benchmarks, we found three layers of signal.
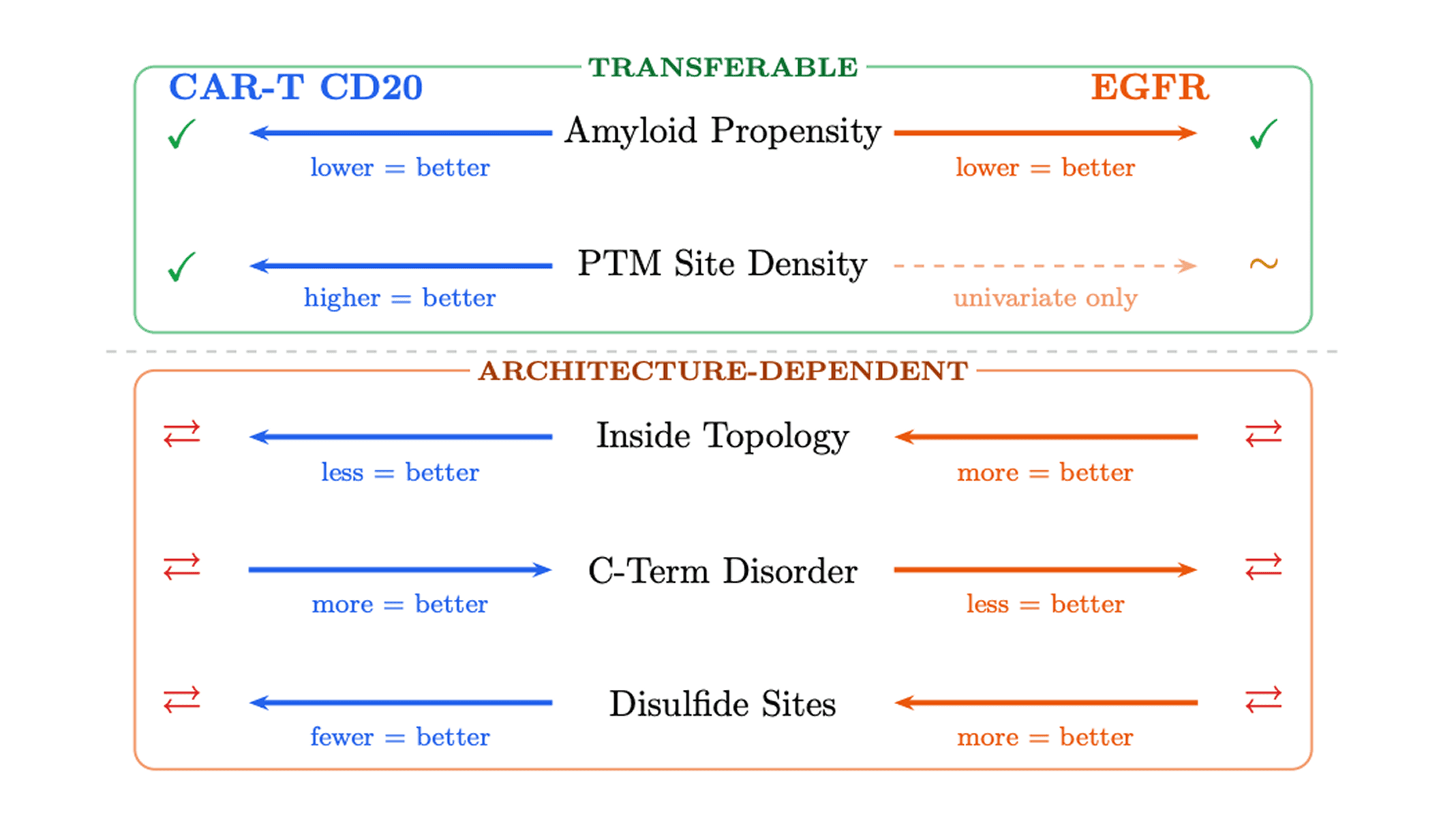
Some signals appear to be transferable. Lower aggregation propensity was the most robust shared association with success across both benchmarks. Predicted PTM-site density also showed a univariate association in both settings, although that result was partly confounded by sequence length in the EGFR dataset.
Some signals are architecture-dependent. Topology-like character, disorder, and disulfide-related features were significant in both datasets, but they flipped direction between contexts. A feature associated with success in a membrane-displayed CAR-T binder could be associated with failure in a standalone EGFR binder, and vice versa.
Some signals are context-specific. In CAR-T, we observed a phosphorylation-related depletion association and a tradeoff between expression and enrichment. In EGFR, low disorder emerged as the dominant binding-associated signal.
One practical result from the study was a retrospective filter analysis in the CAR-T benchmark. Starting from the benchmark’s baseline controlled subset, stacking biology-informed filters increased the enrichment hit rate from 13.8% to 38.6%, a 2.8× lift. This was a retrospective within-benchmark result, not a prospective production model, but it illustrates the potential value of adding a context-aware biological screening layer before synthesis and testing.
The broader takeaway is straightforward: binder evaluation should not rely on one universal scoring logic.
A candidate does not just need to look strong structurally. It also needs to be compatible with how it will be expressed, folded, deployed, and tested. The same sequence descriptor can mean different things depending on whether the binder is part of a CAR construct or a standalone protein.
That is the core idea behind this preprint: binder success depends on deployment context.
We think this matters for anyone building or screening binders computationally. Biology-informed features may help teams not only rank candidates, but also distinguish what kind of failure risk a design carries — aggregation burden, architecture mismatch, expression liability, or loss of functional compatibility in a specific setting.
This paper is a retrospective re-analysis of public benchmark data, and the proposed framework still needs prospective validation. But we believe it points toward a more useful way to think about binder screening: layered, context-aware, and closer to real experimental constraints.
You can read the full preprint here: https://www.biorxiv.org/content/10.64898/2026.04.13.718094v1
Book a 20-minute demo
Sign up free for unlimited Overview runs — summary, sequence-based analysis, homology search. For the full Characterization — PTMs, binding sites, stability variants, construct design — book a demo and we'll run your target live.
