Blog
Orbion Team
Codon Optimization Doesn't Fix Everything
You ordered a codon-optimized gene from your favorite synthesis vendor. The Codon Adaptation Index went from 0.45 to 0.95. E. coli should love this sequence. You transform, induce, and... nothing. No expression. Or worse—the same inclusion body problem you had with the native gene, just with more expensive DNA.
You just learned the hard way that codon optimization solves codon problems. If your expression failure isn't a codon problem—and it usually isn't—optimization won't help.
Key Takeaways
Codon optimization improves expression in only ~30–40% of cases where expression was already failing; it's not a universal fix
Most expression failures are caused by protein-level problems (misfolding, toxicity, PTM requirements) that no codon change can address
Over-optimization can actually reduce expression: eliminating rare codons removes translational pauses that some proteins need for proper folding
mRNA secondary structure near the start codon is often more important than codon usage in the coding region
The most impactful optimization is the first 30–50 nucleotides, not the entire gene
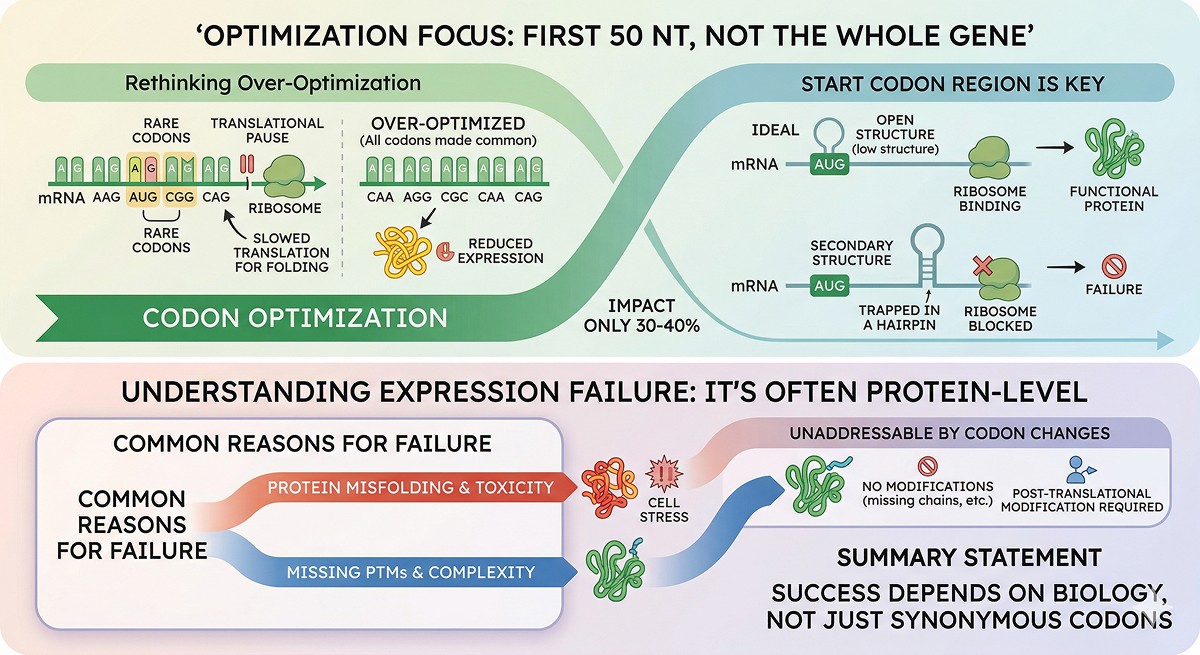
What Codon Optimization Actually Does
The Biology
The genetic code is degenerate: most amino acids are encoded by 2–6 synonymous codons. Different organisms use these codons at different frequencies, and abundant codons are decoded faster because their cognate tRNAs are more available.
Codon optimization replaces codons in your gene of interest with codons preferred by the expression host. For E. coli, this typically means:
Replacing AGG/AGA (rare arginine) with CGU/CGC (common arginine)
Replacing CUA (rare leucine) with CUG (common leucine)
Replacing AUA (rare isoleucine) with AUU (common isoleucine)
Replacing CCC (less common proline) with CCG (common proline)
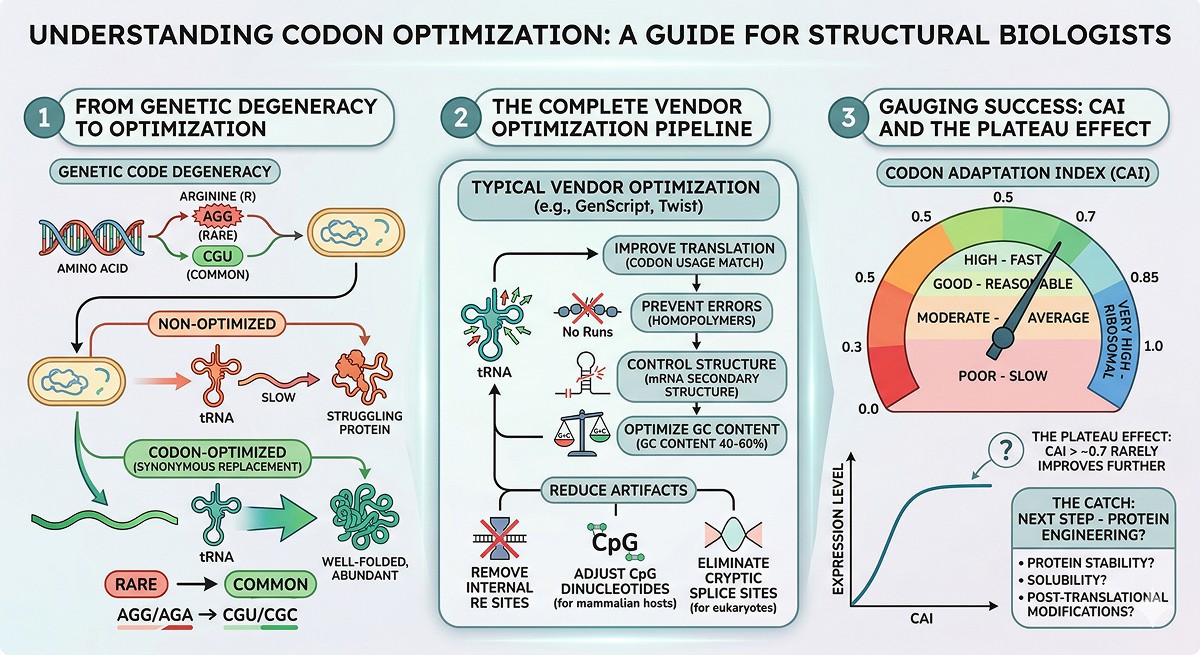
What Vendors Optimize
Modern gene synthesis vendors (GenScript, IDT, Twist, etc.) do more than just swap codons. Typical optimization includes:
Feature | What They Do | Impact |
|---|---|---|
Codon usage | Match host organism frequencies | Moderate (if rare codons were limiting) |
GC content | Normalize to 40–60% | Prevents extreme secondary structures |
Restriction sites | Remove internal RE sites | Enables future cloning |
Homopolymers | Break up poly-A/T/G/C runs | Prevents sequencing/synthesis errors |
mRNA structure | Reduce strong stem-loops | Can improve translation initiation |
CpG dinucleotides | Adjust for host (reduce for mammalian) | Prevents epigenetic silencing in mammals |
Splice sites | Remove cryptic splice signals | Prevents aberrant splicing in eukaryotes |
The Codon Adaptation Index (CAI)
CAI measures how well a gene's codon usage matches the host organism's most highly expressed genes (Sharp & Li, 1987). Scale: 0 to 1.
CAI | Interpretation |
|---|---|
< 0.3 | Very poorly adapted—likely to have slow translation |
0.3–0.5 | Moderate adaptation—typical for average genes |
0.5–0.7 | Good adaptation—should translate reasonably well |
0.7–0.85 | High adaptation—matches highly expressed genes |
> 0.85 | Very high—equivalent to ribosomal protein genes |
The catch: CAI above ~0.7 rarely improves expression further. The gains from optimization plateau, and the remaining expression problems are not codon-related.
Why Optimization Often Doesn't Help
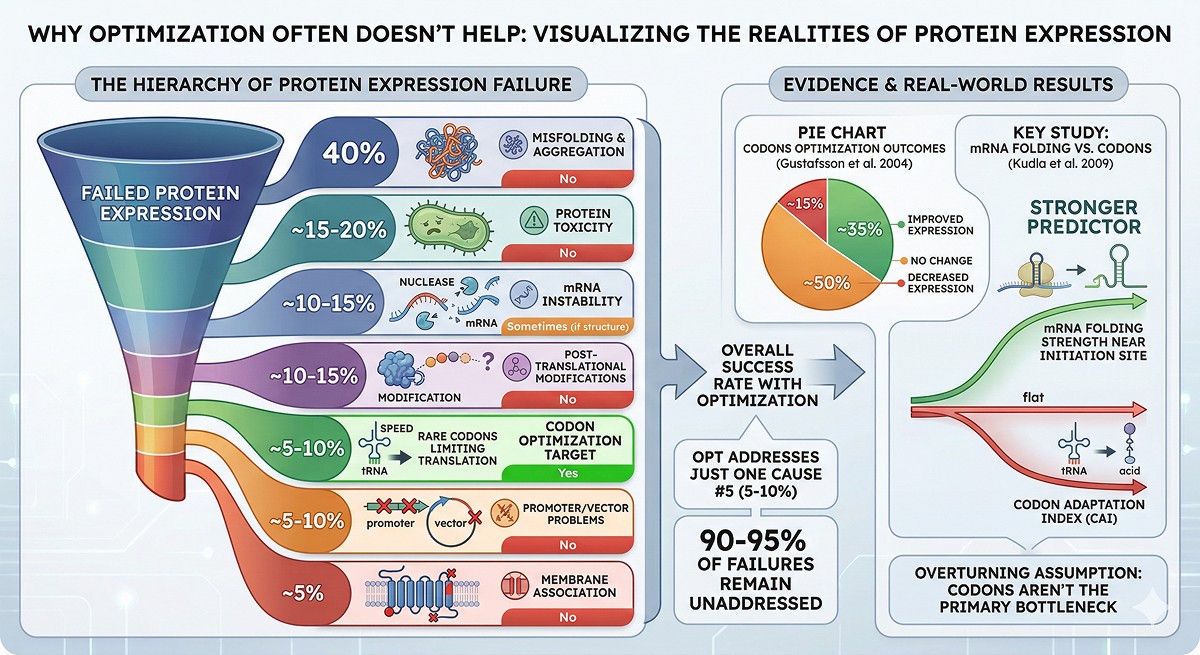
The Hierarchy of Expression Failure
When a protein doesn't express, the cause is almost always one of these, in order of frequency:
Rank | Cause | Frequency | Codon Optimization Helps? |
|---|---|---|---|
1 | Protein misfolding/aggregation | ~40% of failures | No |
2 | Protein toxicity to host | ~15–20% | No |
3 | mRNA instability | ~10–15% | Sometimes (if caused by structure) |
4 | PTM requirements | ~10–15% | No |
5 | Rare codons limiting translation | ~5–10% | Yes |
6 | Promoter/vector problems | ~5–10% | No |
7 | Membrane association | ~5% | No |
Codon optimization addresses cause #5, which accounts for only 5–10% of expression failures. For the other 90–95%, it does nothing.
The Evidence
Gustafsson et al. (2004) reviewed codon optimization outcomes and found:
~30–40% of cases showed improved expression
~50% showed no significant change
~10–20% showed decreased expression
A systematic study by Kudla et al. (2009) demonstrated that for GFP variants in E. coli, mRNA folding near the translation initiation site was a stronger predictor of expression than codon adaptation index—overturning the assumption that rare codons are the primary bottleneck.
When Over-Optimization Hurts
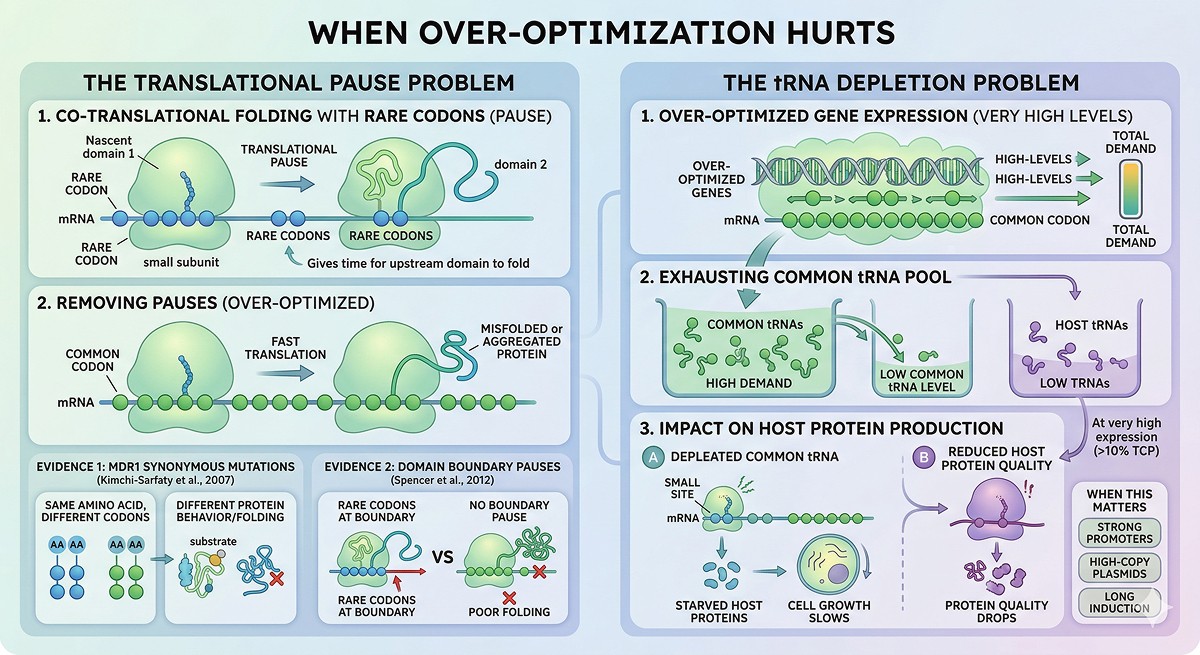
The Translational Pause Problem
Some proteins require slow translation at specific positions for proper co-translational folding. Rare codons create natural pauses that give upstream domains time to fold before downstream domains emerge from the ribosome (Zhang et al., 2009).
What happens when you optimize:
Rare codons → common codons
Translational pauses are removed
Downstream domains emerge before upstream domains finish folding
The protein misfolds or aggregates
Evidence:
Kimchi-Sarfaty et al. (2007) showed that synonymous mutations in the human MDR1 gene altered protein folding and substrate specificity—same amino acid sequence, different codon usage, different protein behavior
Spencer et al. (2012) demonstrated that maintaining rare codons at domain boundaries improved folding of multi-domain proteins in E. coli
Proteins most at risk:
Multi-domain proteins (domain boundaries often use rare codons)
Large proteins (>60 kDa)
Proteins with complex topologies (knotted proteins, β-propellers)
Secreted proteins (signal peptide requires slow initiation)
The tRNA Depletion Problem
If you express a heavily optimized gene at very high levels, you can exhaust the host's tRNA pools for the most common codons. This seems paradoxical, but:
Optimization concentrates demand on a few tRNA species
At very high expression (>10% of total cell protein), those tRNAs become limiting
Other essential host proteins that use the same codons are now starved
Cell growth slows, protein quality drops
When this matters: Only at very high expression levels (strong promoters, high-copy plasmids, long induction). For moderate expression, it's rarely an issue.
The 5' End Matters Most
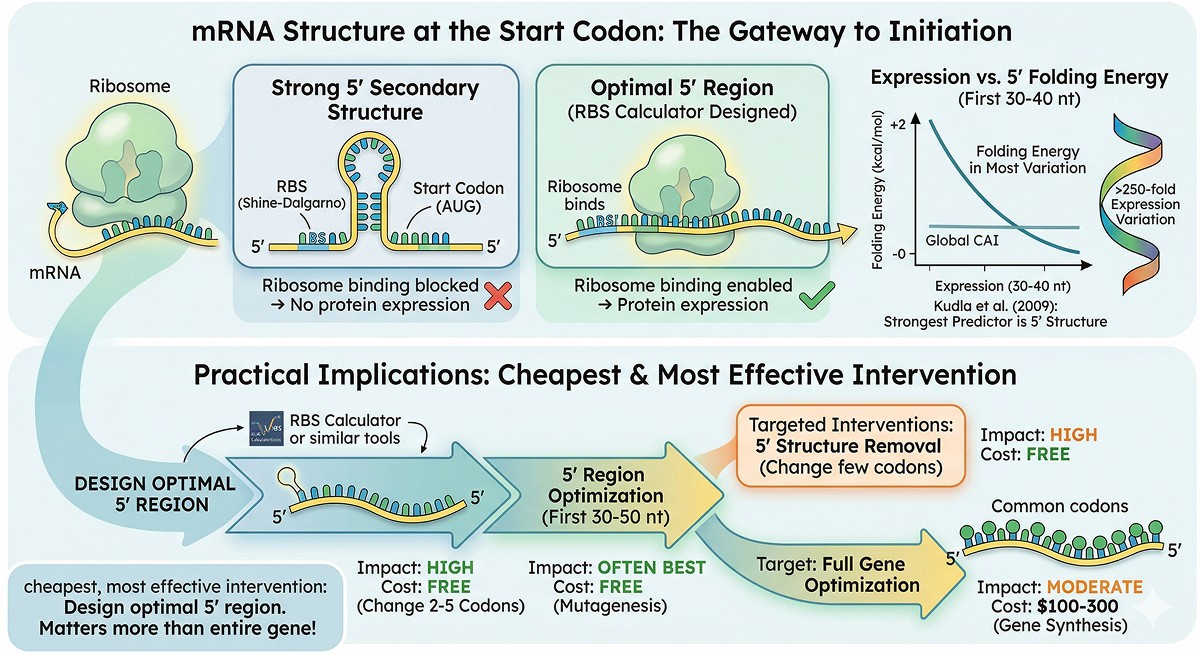
mRNA Structure at the Start Codon
Kudla et al. (2009) showed that the strongest predictor of expression level in E. coli was mRNA secondary structure around the ribosome binding site and start codon, not overall codon usage.
Why:
The ribosome must bind the Shine-Dalgarno sequence and initiate at the AUG
If strong mRNA secondary structure sequesters this region, the ribosome can't bind
No initiation = no protein, regardless of how optimized the downstream codons are
The numbers:
Expression varied >250-fold across GFP variants with identical amino acid sequences
mRNA folding energy in the first 30–40 nucleotides explained most of this variation
Global CAI explained almost none
Practical Implications
Optimization Target | Impact on Expression | Cost |
|---|---|---|
Full gene codon optimization | Moderate (if rare codons were a problem) | $100–300 (gene synthesis) |
5' region optimization only (first 30–50 nt) | Often equal to or better than full optimization | Free (change a few codons by mutagenesis) |
RBS calculator (design optimal ribosome binding site) | High for E. coli expression | Free (online tools) |
Removing 5' mRNA structure | High | Free (change 2–5 codons) |
The cheapest, most effective intervention: Use the Salis Lab RBS Calculator or similar tools to design an optimal 5' region. This often matters more than optimizing the entire gene.
What to Do Instead (or in Addition to) Codon Optimization
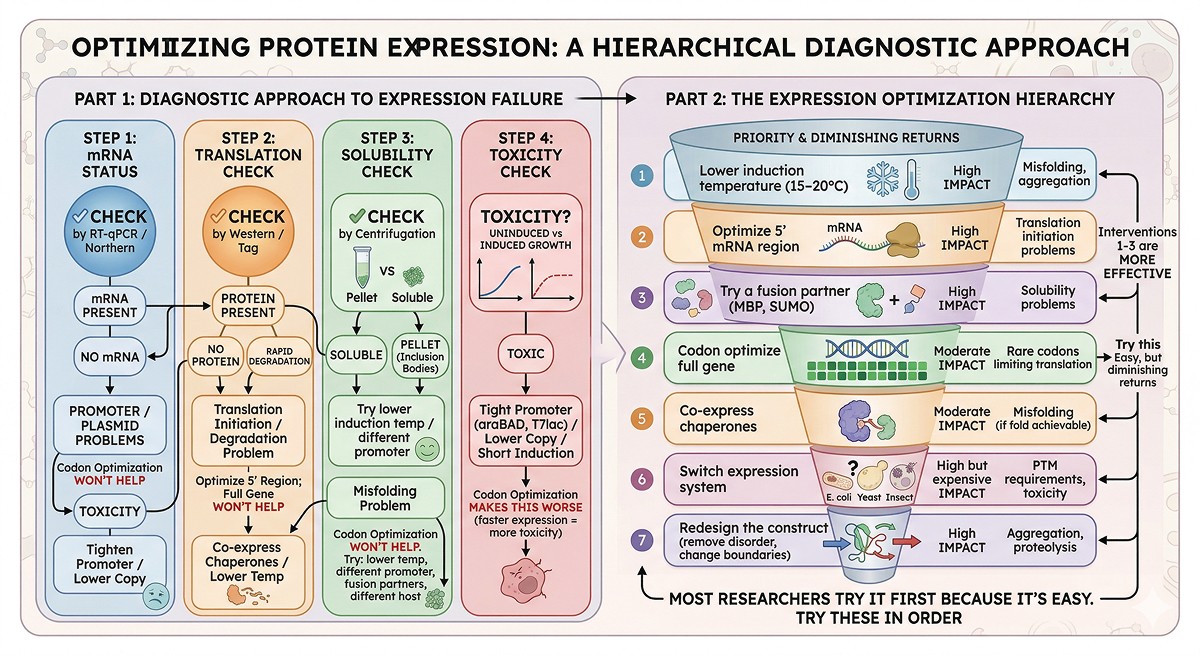
A Diagnostic Approach to Expression Failure
Step 1: Is the mRNA being made?
Check by RT-qPCR or Northern blot
If no mRNA → promoter problem, plasmid problem, or toxicity
Codon optimization won't help
Step 2: Is the protein being translated?
Check by Western blot (if antibody available) or by expressing with a small tag
If mRNA present but no protein → translation initiation problem or rapid degradation
Optimize the 5' region; codon optimization of the full gene is less likely to help
Step 3: Is the protein soluble?
Check by centrifugation: soluble fraction vs pellet (inclusion bodies)
If protein is in the pellet → misfolding problem
Codon optimization won't help. Try: lower temperature, different promoter, fusion partners, different host.
Step 4: Is the protein toxic?
Check: does the uninduced culture grow normally? Does growth halt upon induction?
If toxic → tight promoter (araBAD, T7lac), lower copy number, short induction times
Codon optimization makes this worse (faster expression = more toxicity)
The Expression Optimization Hierarchy
Try these in order—each has diminishing returns:
Priority | Intervention | Expected Impact | When It Helps |
|---|---|---|---|
1 | Lower induction temperature (15–20°C) | High | Misfolding, aggregation |
2 | Optimize 5' mRNA region | High | Translation initiation problems |
3 | Try a fusion partner (MBP, SUMO) | High | Solubility problems |
4 | Codon optimize full gene | Moderate | Rare codons limiting translation |
5 | Co-express chaperones | Moderate | Misfolding (if the fold is achievable) |
6 | Switch expression system | High but expensive | PTM requirements, toxicity |
7 | Redesign the construct (remove disorder, change boundaries) | High | Aggregation, proteolysis |
Codon optimization is priority #4. Most researchers try it first because it's easy. But interventions 1–3 are more frequently effective.
When Codon Optimization IS the Right Answer
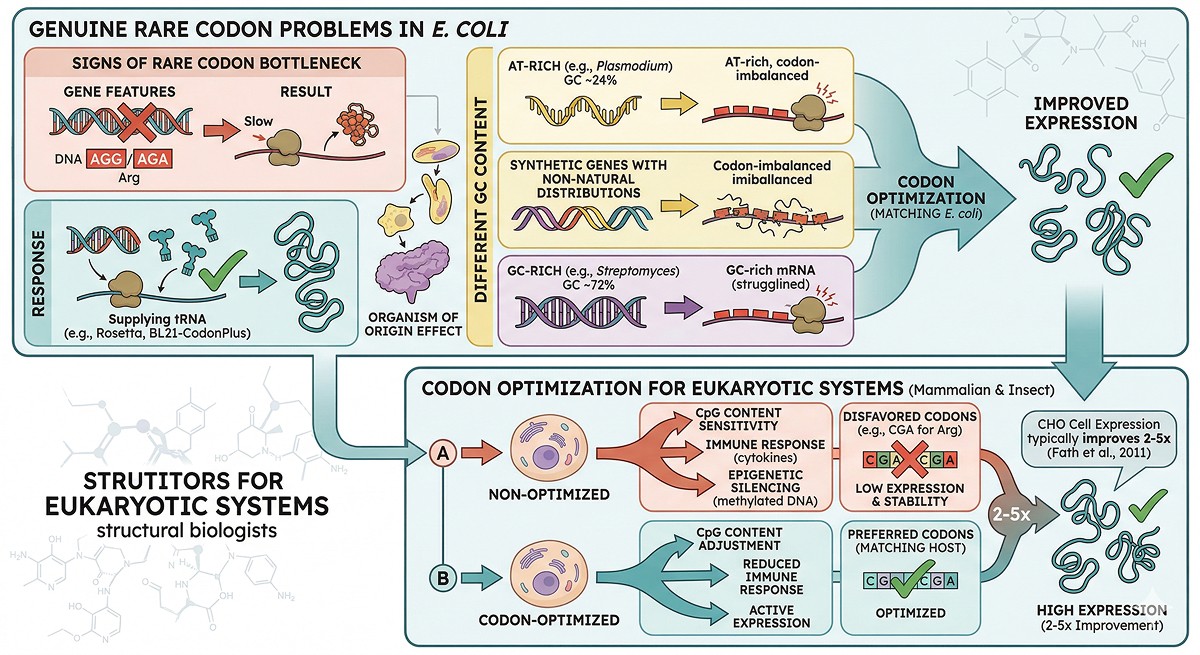
Genuine Rare Codon Problems
Some expression failures are genuinely caused by rare codons:
Signs that rare codons are your problem:
The gene has clusters of rare codons (e.g., 3+ AGG/AGA in a row)
Expression improves with BL21-CodonPlus or Rosetta strains (which supply extra tRNAs)
The organism of origin has very different codon usage from E. coli (e.g., AT-rich genomes from Plasmodium, or GC-rich genes from Streptomyces)
When optimization clearly helps:
Genes from AT-rich organisms (Plasmodium: average GC ~24% vs E. coli ~51%)
Genes from GC-rich organisms (Streptomyces: average GC ~72%)
Synthetic genes with non-natural codon distributions
Genes with extreme amino acid compositions (Arg/Ile/Leu-rich from organisms with different codon preferences)
Eukaryotic Expression Systems
Codon optimization matters more for mammalian and insect cell expression than for E. coli:
Mammalian cells are more sensitive to CpG content (can trigger immune responses or epigenetic silencing)
Some codons are strongly disfavored in mammalian systems (e.g., CGA for arginine)
Codon optimization for CHO cells typically improves expression 2–5x (Fath et al., 2011)
Beyond Codons: What Else to Optimize in Your Gene
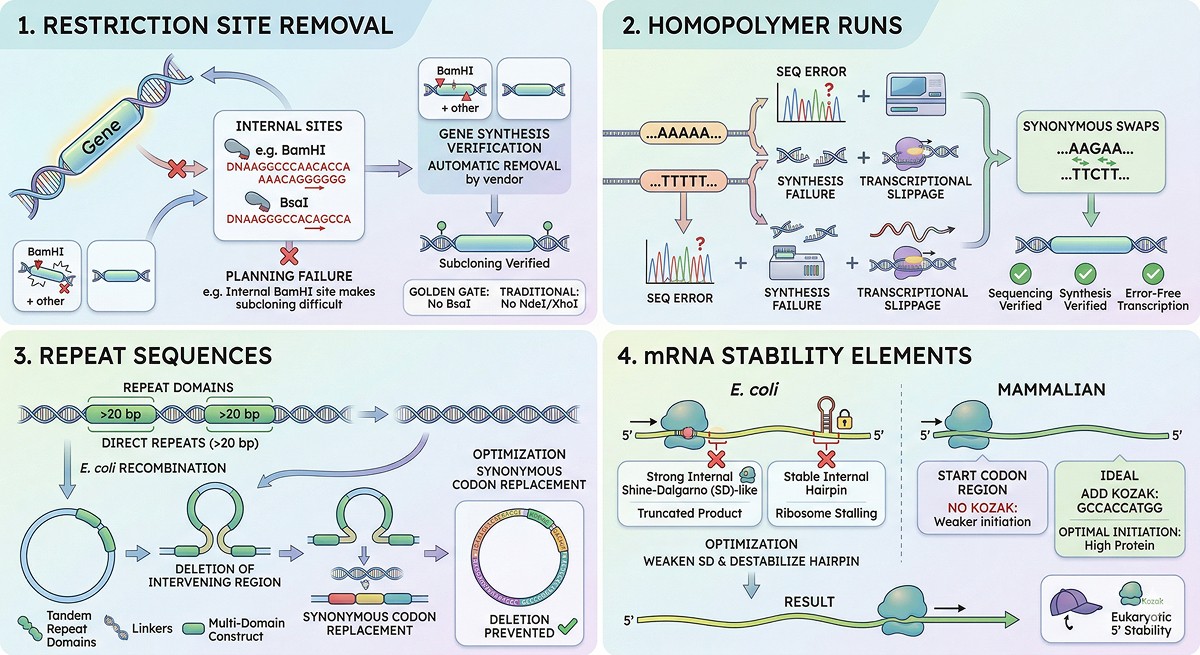
Restriction Site Removal
If you plan to subclone, having internal restriction sites in your gene is a practical nightmare. Gene synthesis vendors handle this automatically, but verify:
No BamHI, EcoRI, NdeI, XhoI sites in the coding region (or whatever your cloning strategy requires)
No BsaI sites if using Golden Gate assembly
Homopolymer Runs
Stretches of >5 identical nucleotides can cause:
Sequencing errors (polymerase slippage)
Synthesis failures (difficult to assemble)
Transcriptional errors in vivo
Vendors break these up by synonymous codon swaps.
Repeat Sequences
Direct repeats >20 bp can cause recombination in E. coli, deleting the region between repeats. This is particularly relevant for:
Proteins with tandem repeat domains
Codon-optimized genes where the limited codon palette creates accidental repeats
Multi-domain constructs with repeated linker sequences
mRNA Stability Elements
Beyond the 5' region:
Avoid strong internal Shine-Dalgarno-like sequences (can cause internal translation initiation → truncated products)
Minimize very stable hairpins in the coding region (can stall the ribosome)
For mammalian expression, consider adding a Kozak sequence (GCCACCATGG) around the start codon
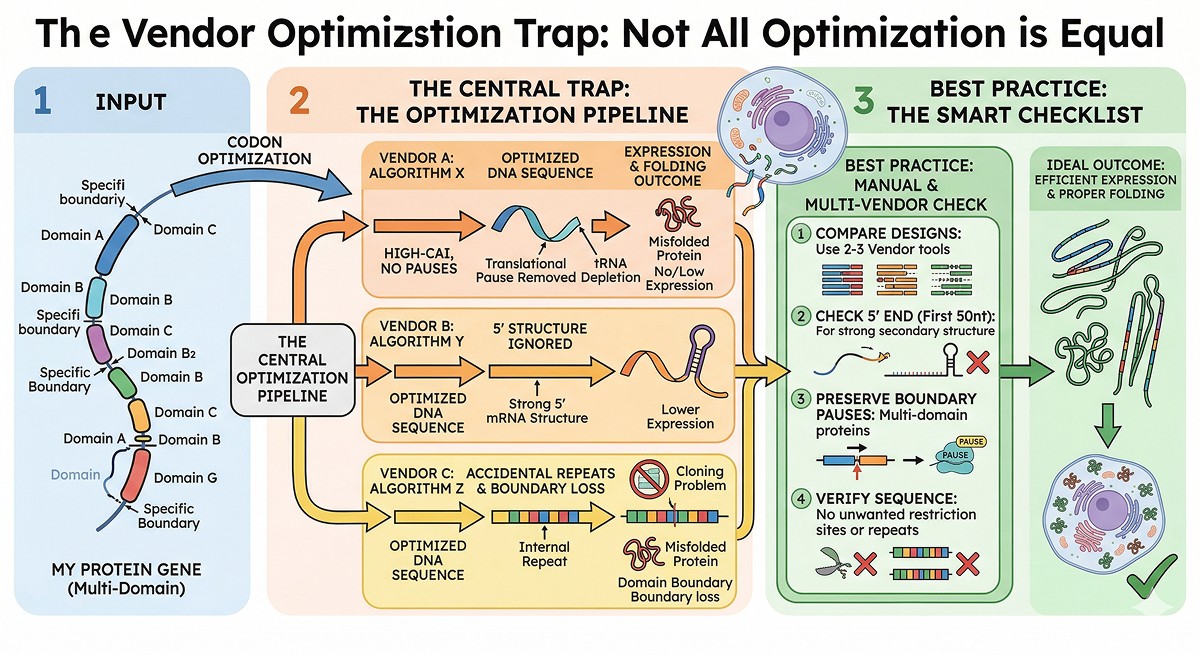
The Vendor Optimization Trap
Not All "Optimization" Is Equal
Different vendors use different algorithms. Testing the same protein with three vendors will give three different DNA sequences—and potentially different expression levels.
What to watch for:
Some algorithms over-optimize, removing ALL rare codons (risks: translational pause removal, tRNA depletion)
Some don't adequately address mRNA structure at the 5' end
Some create sequences with internal repeat regions that cause cloning problems
Few consider co-translational folding requirements
Best practice:
Use 2–3 vendor tools and compare the designs
Manually check the first 50 nucleotides for strong secondary structure
If your protein is multi-domain, consider preserving rare codons at domain boundaries
Verify that no unwanted restriction sites or repeats were introduced
The Bottom Line
Expression Problem | Will Codon Optimization Help? | Better Intervention |
|---|---|---|
No expression at all | Maybe (if rare codons limit translation) | Check mRNA levels first; optimize 5' region |
Expression but insoluble | No | Lower temperature, fusion partner, construct redesign |
Low yield | Maybe (if codon-limited) | CodonPlus strain test; 5' optimization |
Protein is toxic | No (may make it worse) | Tight promoter, low-copy vector |
Needs PTMs | No | Switch expression system |
Protein degrades rapidly | No | Protease-deficient strain, remove disordered regions |
Gene from very different organism | Yes (likely codon-limited) | Optimize + consider codon harmony |
The core message: Codon optimization is one tool in the expression toolbox, not a magic fix. Diagnose the actual problem before spending money on gene synthesis. In most cases, the failure has nothing to do with codons.
Smarter Construct Design with Orbion
Orbion's Construct Design module handles codon optimization as part of a broader strategy—not in isolation. The system performs organism-specific codon optimization while avoiding restriction sites, homopolymers, and GC extremes. But more importantly, it integrates codon optimization with construct boundary design (removing disordered regions before optimizing) and tag/fusion partner selection (addressing the folding problem that codons can't fix).
AstraSUIT predicts expression system suitability upfront—flagging proteins that need glycosylation, membrane insertion, or specific cofactors that no amount of codon optimization will provide. The Bench module then generates expression protocols matched to the construct design, so optimization happens at every level: sequence, construct, expression conditions, and purification strategy.
References
Sharp PM & Li WH. (1987). The codon adaptation index—a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Research, 15(3):1281-1295. PMC340524
Gustafsson C, et al. (2004). Codon bias and heterologous protein expression. Trends in Biotechnology, 22(7):346-353. Link
Kudla G, et al. (2009). Coding-sequence determinants of gene expression in Escherichia coli. Science, 324(5924):255-258. Link
Kimchi-Sarfaty C, et al. (2007). A "silent" polymorphism in the MDR1 gene changes substrate specificity. Science, 315(5811):525-528. Link
Zhang G, et al. (2009). Transient ribosomal attenuation coordinates protein synthesis and co-translational folding. Nature Structural & Molecular Biology, 16:274-280. Link
Spencer PS, et al. (2012). Silent substitutions predictably alter translation elongation rates and protein folding efficiencies. Journal of Molecular Biology, 422(3):328-335. Link
Fath S, et al. (2011). Multiparameter RNA and codon optimization: a standardized tool to assess and enhance autologous mammalian gene expression. PLoS ONE, 6(3):e17596. Link
Plotkin JB & Kudla G. (2011). Synonymous but not the same: the causes and consequences of codon bias. Nature Reviews Genetics, 12:32-42. Link
Chaney JL & Clark PL. (2015). Roles for synonymous codon usage in protein biogenesis. Annual Review of Biophysics, 44:143-166. Link
Angov E, et al. (2008). Adjustment of codon usage frequencies by codon harmonization improves protein expression and folding. Methods in Molecular Biology, 459:1-16. Link
