Blog
From Single Mutants to Combinatorial Libraries: Scaling Protein Engineering
Mar 27, 2026

You identified eight single mutations that each improve thermostability by 2–5°C. Beautiful, consistent ΔTm data from differential scanning fluorimetry. You combine all eight into one construct, expecting +20–30°C. You get –3°C. Less stable than wild-type. The eight "beneficial" mutations, when combined, produce a protein that aggregates during purification.
Welcome to epistasis—the reason that protein engineering doesn't scale linearly and why scaling from single mutants to successful combinatorial variants is one of the hardest problems in the field.
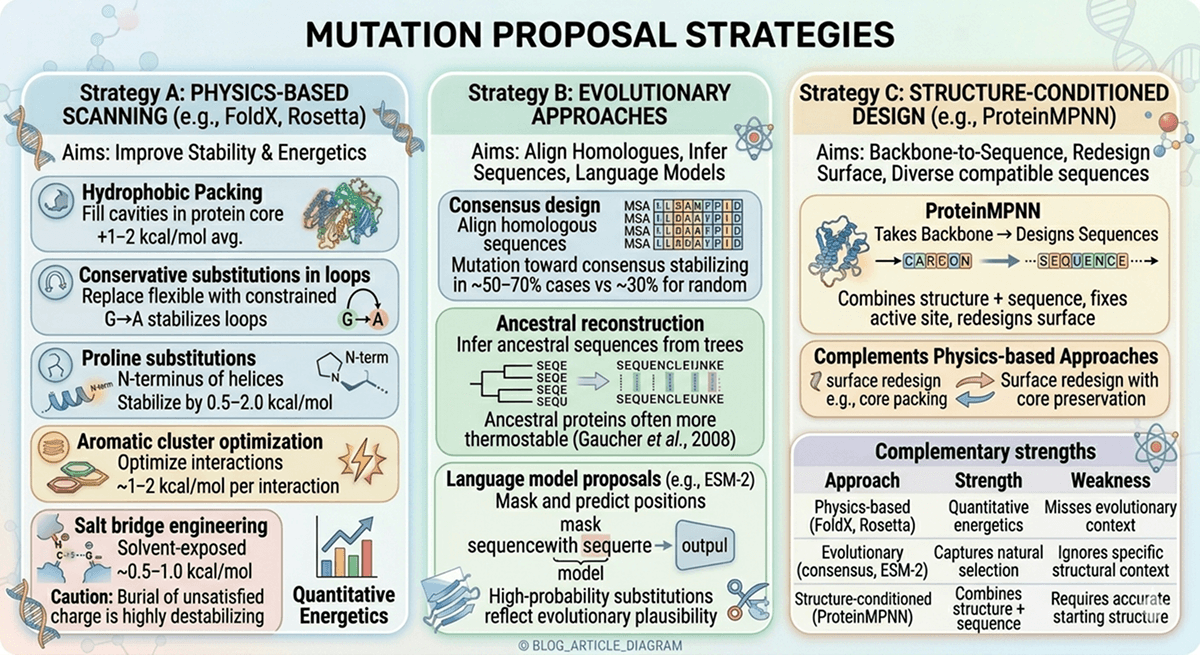
Key Takeaways
Beneficial single mutations rarely add up: Epistasis means that the effect of combining mutations is unpredictable from individual effects alone
The combinatorial space is astronomically large: Even 10 positions with 2 options each create 1,024 variants—far more than most labs can screen
Smart library design beats brute-force screening: Structure-guided clustering, computational pre-screening, and evolutionary information dramatically reduce the search space
Multi-objective optimization is essential: Stability, activity, expression, and aggregation resistance often trade off against each other
Genetic algorithms and staged evaluation can navigate fitness landscapes efficiently without exhaustive screening
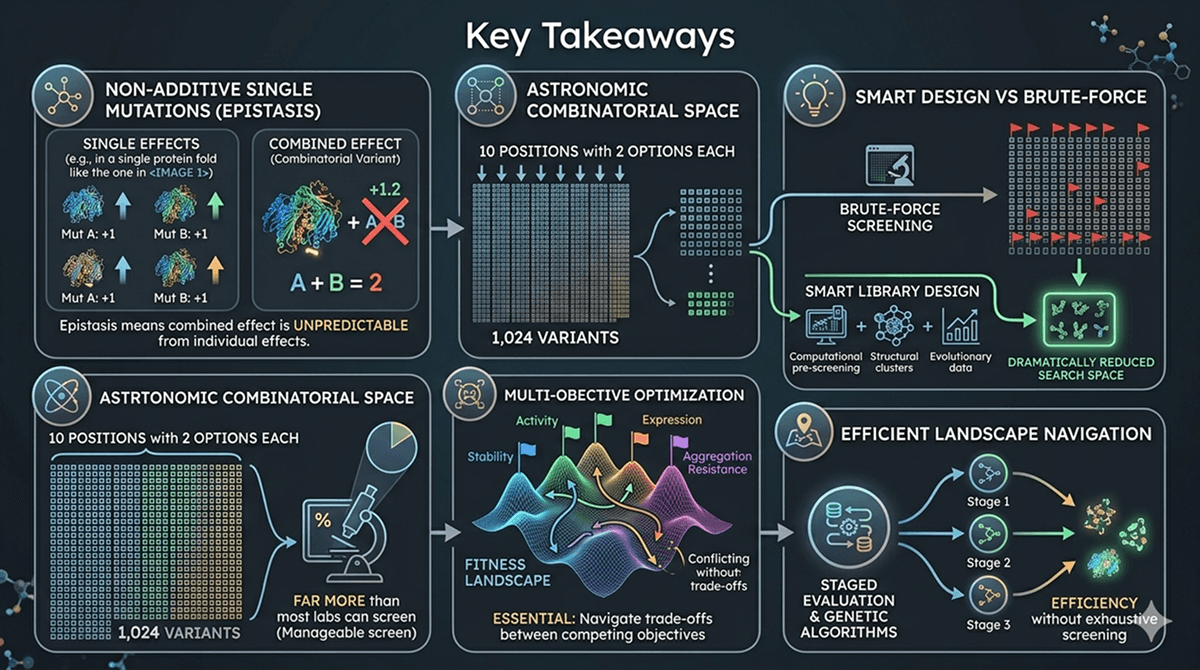
Why Single Mutant Data Doesn't Add Up
The Additivity Assumption
The simplest model for combining mutations assumes additivity:
ΔΔG(A+B) = ΔΔG(A) + ΔΔG(B)
If mutation A stabilizes by 1.5 kcal/mol and mutation B by 2.0 kcal/mol, the double mutant should stabilize by 3.5 kcal/mol.
This is wrong more often than it's right.
What Actually Happens: Epistasis
Epistasis—the nonadditive interaction between mutations—means that the effect of one mutation depends on what other mutations are present (Starr & Thornton, 2016).
Types of epistasis:
Type | Definition | Example | Consequence |
|---|---|---|---|
Magnitude | Combined effect is larger or smaller than expected | A (+2°C) + B (+3°C) = AB (+7°C) or AB (+3°C) | Under- or over-estimation of combined benefit |
Sign | One mutation changes from beneficial to harmful | A (+2°C) alone, but A (–1°C) in presence of B | "Beneficial" single mutants can be harmful in combination |
Reciprocal sign | Both mutations beneficial alone, harmful together | A (+2°C), B (+3°C), AB (–3°C) | The nightmare scenario—your best singles are incompatible |
Specific | Effect depends on exact genetic background | A helps with B but hurts with C | Order and context matter for every combination |
How Common Is Epistasis?
Very common. Studies of systematic mutation combinations show:
~30–50% of mutation pairs show significant epistasis
Sign epistasis (beneficial → harmful) occurs in ~10–25% of pairs
Reciprocal sign epistasis occurs in ~5–15% of pairs
Epistasis is more common for mutations that are structurally close to each other
The practical implication: If you have 8 beneficial singles, the probability that all 8 are compatible is roughly (0.75)^28 ≈ 0.03, assuming ~25% pairwise epistasis across 28 pairs. In other words, there's about a 3% chance the full combination works as expected.
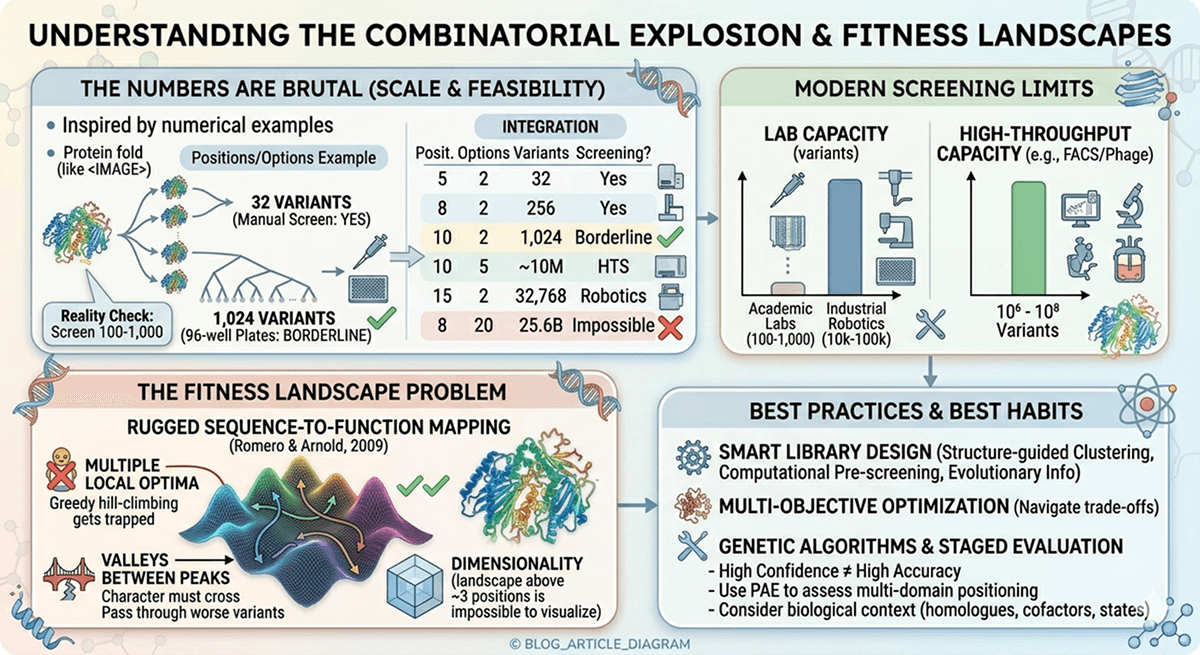
The Combinatorial Explosion
The Numbers Are Brutal
Consider a modest protein engineering campaign:
Positions | Options per Position | Total Variants | Screening Feasible? |
|---|---|---|---|
5 | 2 (WT + mutant) | 32 | Yes (manually) |
8 | 2 | 256 | Yes (96-well plates) |
10 | 2 | 1,024 | Borderline |
10 | 5 | ~10 million | No (need HTS) |
15 | 2 | 32,768 | Need robotics |
8 | 20 (all amino acids) | 25.6 billion | Impossible |
Reality check: Most academic labs can screen 100–1,000 variants per campaign. Industrial labs with robotics: 10,000–100,000. Even directed evolution with FACS can screen ~10^6–10^8. But 20^8 = 25.6 billion. You'll never screen everything.
The Fitness Landscape Problem
Protein fitness landscapes—the mapping from sequence to function—are rugged (Romero & Arnold, 2009). This means:
Multiple local optima: Greedy hill-climbing gets trapped
Valleys between peaks: Getting from one good variant to a better one may require passing through worse variants
Dimensionality: Every mutation adds a dimension; the landscape is impossible to visualize above ~3 positions
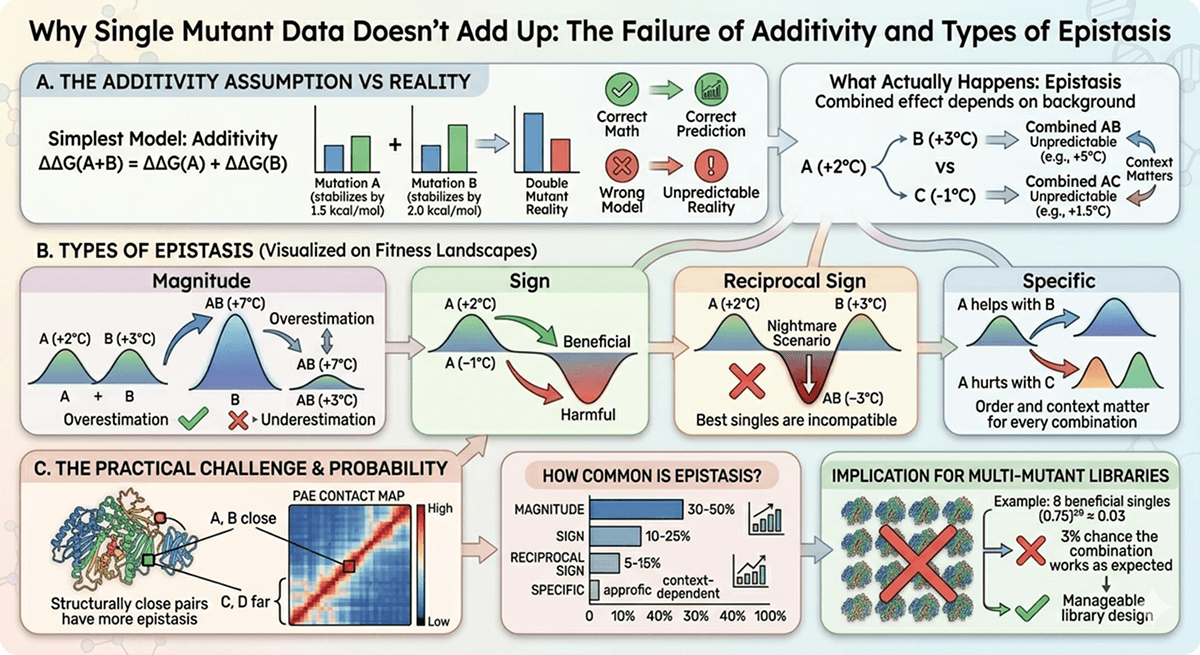
Smart Library Design Strategies
Strategy 1: Recombination of Beneficial Singles
The approach: Start with N beneficial single mutations. Instead of making all 2^N combinations, use structure-guided selection.
Step 1: Cluster by structural region
Group your beneficial mutations by their location in the structure:
Core packing mutations (buried residues)
Surface charge mutations (exposed residues)
Interface mutations (if relevant)
Loop/turn mutations
Step 2: Combine across clusters, not within
Mutations in different structural regions are less likely to show epistasis than mutations in the same region. Combine one mutation from each cluster.
Example:
Cluster A (core): V45I, L89M (pick one)
Cluster B (surface): E120K, D155N (pick one)
Cluster C (loop): G201A (only one)
This gives 2 × 2 × 1 = 4 combinations instead of 2^5 = 32. Each combination contains structurally independent mutations.
Strategy 2: Computational Pre-Screening
The approach: Use computational methods to evaluate combinations before making them in the lab.
Tier 1 (fast, approximate):
Additive ΔΔG estimates (quick but misses epistasis)
Rosetta combinatorial scoring (Kellogg et al., 2011)
Machine learning predictions (ΔTm, ΔΔG from sequence features)
Tier 2 (slower, better):
FoldX BuildModel with full side-chain repacking (Schymkowitz et al., 2005)
Rosetta with full backbone relaxation
Structure prediction of top variants (AlphaFold2)
Tier 3 (expensive, most accurate):
Molecular dynamics of candidate combinations
Free energy perturbation calculations
The practical pipeline:
Score all pairwise combinations computationally (Tier 1): ~minutes
Filter to top 20% of combinations
Re-score with Tier 2 methods: ~hours
Select top 50–100 for experimental testing
Strategy 3: Directed Evolution with Smart Seeding
The approach: Instead of starting directed evolution from wild-type, seed the library with your best computational predictions.
Advantages:
Starts from a higher fitness baseline
Explores epistatic interactions through random recombination
Selection pressure handles what computation can't predict
Library types:
Library Type | Best For | Typical Size | Method |
|---|---|---|---|
Error-prone PCR | Exploring nearby sequence space | 10^3–10^5 | Random mutagenesis |
Site-saturation (NNK) | Optimizing specific positions | 20^N per position | Focused diversity |
DNA shuffling | Recombining beneficial blocks | 10^4–10^6 | Homologous recombination |
SCHEMA | Recombining structural domains | 10^2–10^4 | Structure-guided recombination |
Machine learning-guided | Navigating fitness landscapes | 10^2–10^3 (focused) | Bayesian optimization |

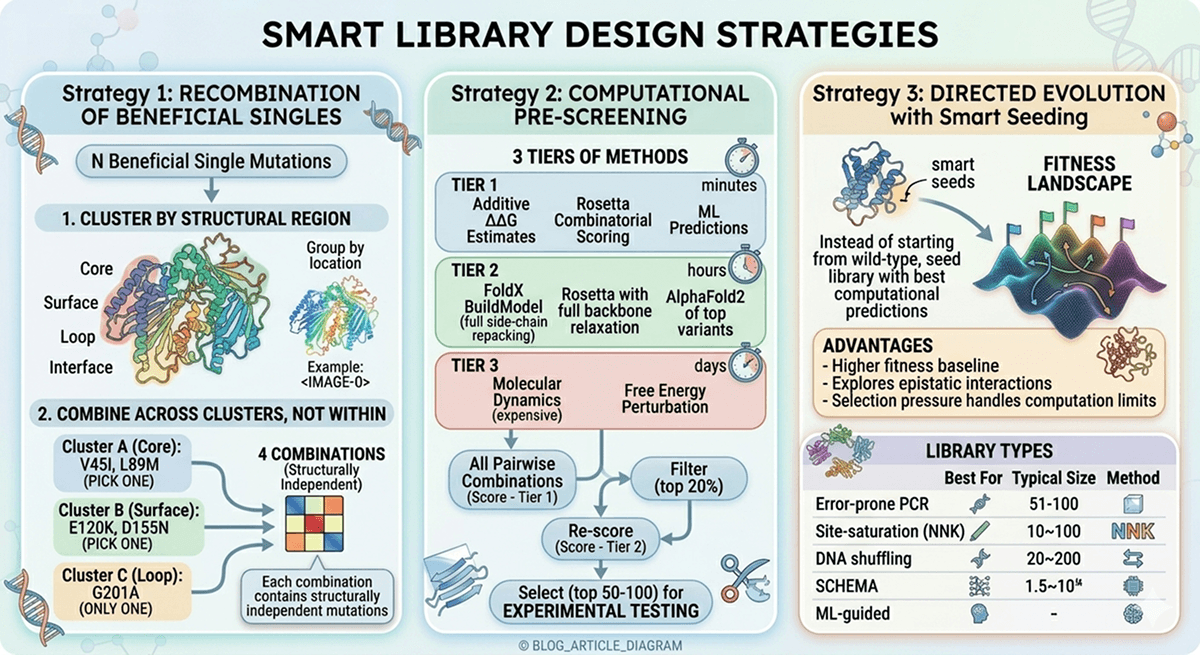
Mutation Proposal Strategies
Physics-Based Approaches
Physics-based scanning identifies mutations likely to improve stability based on structural principles:
Hydrophobic packing improvements:
Fill small cavities in the protein core
Replace small hydrophobics with larger ones where space permits (A→V, V→I, V→L)
Cavity-filling mutations improve stability by ~1–2 kcal/mol on average
Conservative substitutions in loops:
Replace flexible residues with conformationally constrained ones (G→A in loops)
Proline substitutions at the N-terminus of helices
Aromatic cluster optimization:
Introduce or optimize aromatic-aromatic interactions
Phenylalanine/tyrosine stacking contributes ~1–2 kcal/mol per interaction
Salt bridge engineering:
Surface salt bridges contribute ~0.5–1.0 kcal/mol
Most effective when both residues are solvent-exposed
Caution: burial of unsatisfied charges is highly destabilizing
Evolutionary Approaches
Consensus design:
Align homologous sequences
At each position, identify the most frequent amino acid
Mutations toward consensus are stabilizing more often than not
Consensus mutations stabilize in ~50–70% of cases vs ~30% for random mutations
Ancestral reconstruction:
Infer ancestral sequences from phylogenetic trees
Ancestral proteins are often more thermostable (Gaucher et al., 2008)
Mutations toward ancestral sequence can be combinatorially favorable
Language model proposals (ESM-2):
Mask each position and ask what the model predicts
High-probability substitutions reflect evolutionary plausibility
These mutations are less likely to disrupt fold or function
Structure-Conditioned Design
ProteinMPNN (Dauparas et al., 2022):
Takes a backbone structure as input
Designs sequences that would fold into that structure
Can fix certain positions (active site) while redesigning others
Generates diverse sequences that are structurally compatible
Particularly powerful for surface redesign while preserving core
How ProteinMPNN complements physics-based approaches:
Approach | Strength | Weakness |
|---|---|---|
Physics-based (FoldX, Rosetta) | Quantitative energetics | Misses evolutionary context |
Evolutionary (consensus, ESM-2) | Captures natural selection | Ignores specific structural context |
Structure-conditioned (ProteinMPNN) | Combines structure + sequence | Requires accurate starting structure |
Multi-Objective Optimization
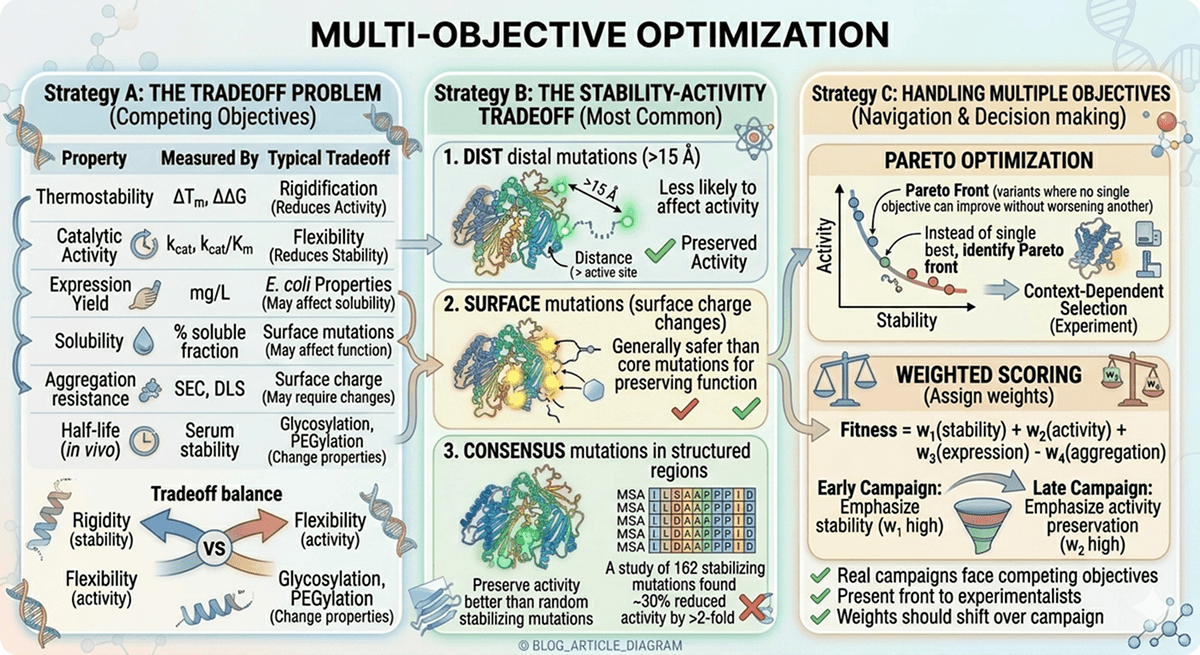
The Tradeoff Problem
Protein engineering rarely optimizes one property. Real campaigns face competing objectives:
Property | Measured By | Typical Tradeoff |
|---|---|---|
Thermostability | ΔTm, ΔΔG | May reduce activity (rigidification) |
Catalytic activity | kcat, kcat/Km | May reduce stability (flexibility needed) |
Expression yield | mg/L culture | May select for E. coli-preferred properties |
Solubility | % soluble fraction | Surface mutations may affect function |
Aggregation resistance | SEC, DLS | May require surface charge changes |
Half-life (in vivo) | Serum stability | Glycosylation, PEGylation change properties |
The Stability-Activity Tradeoff
The most common tradeoff in protein engineering. Stabilizing mutations often rigidify the protein, which can reduce catalytic activity:
Distal mutations (>15 Å from active site) are less likely to affect activity
Surface mutations generally safer than core mutations for preserving function
Consensus mutations in structured regions tend to preserve activity better than random stabilizing mutations
A study of 162 stabilizing mutations found that ~30% reduced activity by >2-fold
Handling Multiple Objectives
Pareto optimization:
Instead of a single "best" variant, identify the Pareto front—variants where no single objective can improve without worsening another
Present the front to experimentalists for context-dependent selection
Weighted scoring:
Assign weights to each objective based on project priorities
Fitness = w₁(stability) + w₂(activity) + w₃(expression) - w₄(aggregation)
Weights should shift over the campaign (early: emphasize stability; late: emphasize activity preservation)
Genetic Algorithms for Variant Optimization
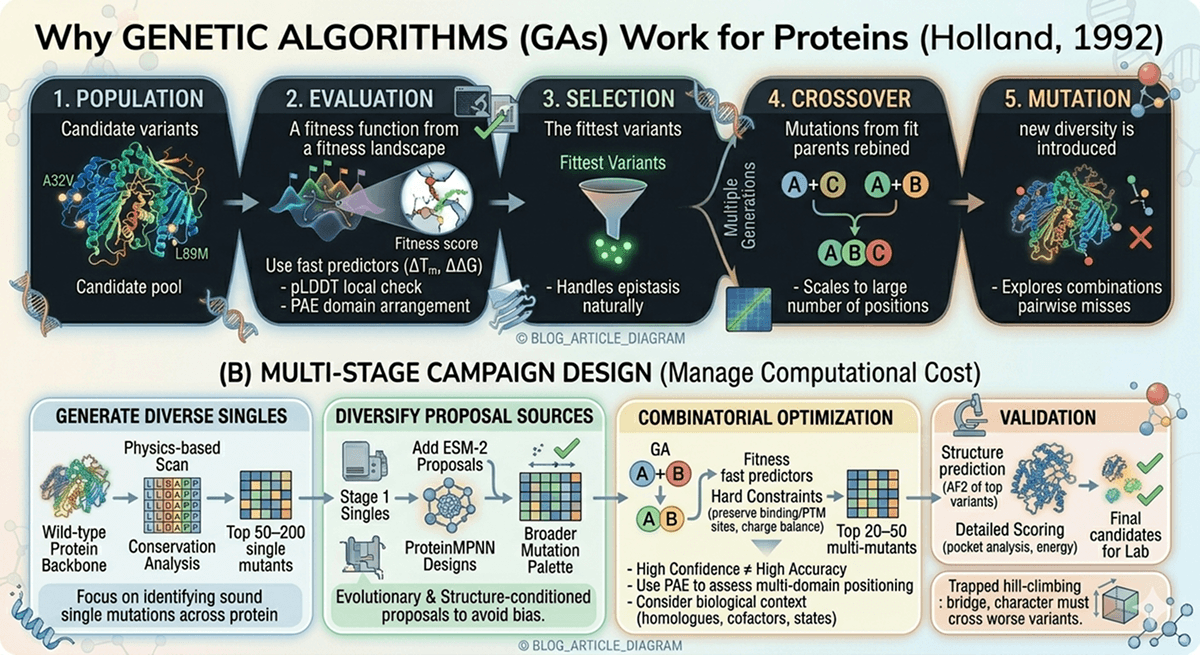
Why Genetic Algorithms Work for Proteins
Genetic algorithms (GAs) mimic natural evolution to navigate combinatorial landscapes (Holland, 1992):
Population: Start with a set of candidate variants
Evaluation: Score each variant for fitness (stability, activity, etc.)
Selection: Keep the fittest variants
Crossover: Recombine mutations from fit parents
Mutation: Introduce new diversity
Repeat: Run for multiple generations
Why this works for proteins:
Explores combinations that simple pairwise screening misses
Naturally handles epistasis through selection pressure
Scales to large numbers of positions
Can incorporate multiple objectives via fitness function design
Multi-Stage Campaign Design
The most effective campaigns use staged evaluation to manage computational cost:
Stage | Goal | Methods | Output |
|---|---|---|---|
1 | Generate diverse singles | Physics-based scan, conservation analysis | Top 50–200 single mutants |
2 | Diversify proposal sources | Add ESM-2 proposals, ProteinMPNN designs | Broader mutation palette |
3 | Combinatorial optimization | Genetic algorithm with fast fitness evaluation | Top 20–50 multi-mutants |
4 | Validation | Structure prediction, detailed scoring | Final candidates for lab |
Stage 1 focuses on identifying structurally sound single mutations across the protein.
Stage 2 adds evolutionary and structure-conditioned proposals to avoid bias from physics alone.
Stage 3 is the key innovation: use a genetic algorithm to recombine mutations from Stages 1-2, scoring each combination with fast predictors (ΔTm, ΔΔG, disorder) and applying hard constraints (preserve binding sites, PTM sites, charge balance).
Stage 4 validates the top candidates from Stage 3 with expensive methods (AlphaFold2 structure prediction, RMSD to wild-type, pocket analysis).
Real-World Success Stories
Industrial Enzyme Stabilization
Arnold and colleagues demonstrated that directed evolution can dramatically improve enzyme thermostability—in some cases by >30°C—but required thousands of variants screened over multiple rounds.
Key lesson: Combining rational pre-selection with directed evolution reduced the number of rounds needed from ~5–10 to 2–3.
Therapeutic Antibody Engineering
Antibody optimization campaigns routinely combine:
CDR mutations for affinity maturation
Framework mutations for stability
Fc mutations for effector function
Successful campaigns report that computational pre-screening reduced experimental testing by 10-100x while maintaining the same hit rate.
Ancestral Reconstruction + Rational Design
Risso et al. (2013) combined ancestral reconstruction with rational design to create a β-lactamase with ΔTm > +30°C while retaining catalytic activity. The key was using the ancestral sequence as a starting point for combinatorial optimization.
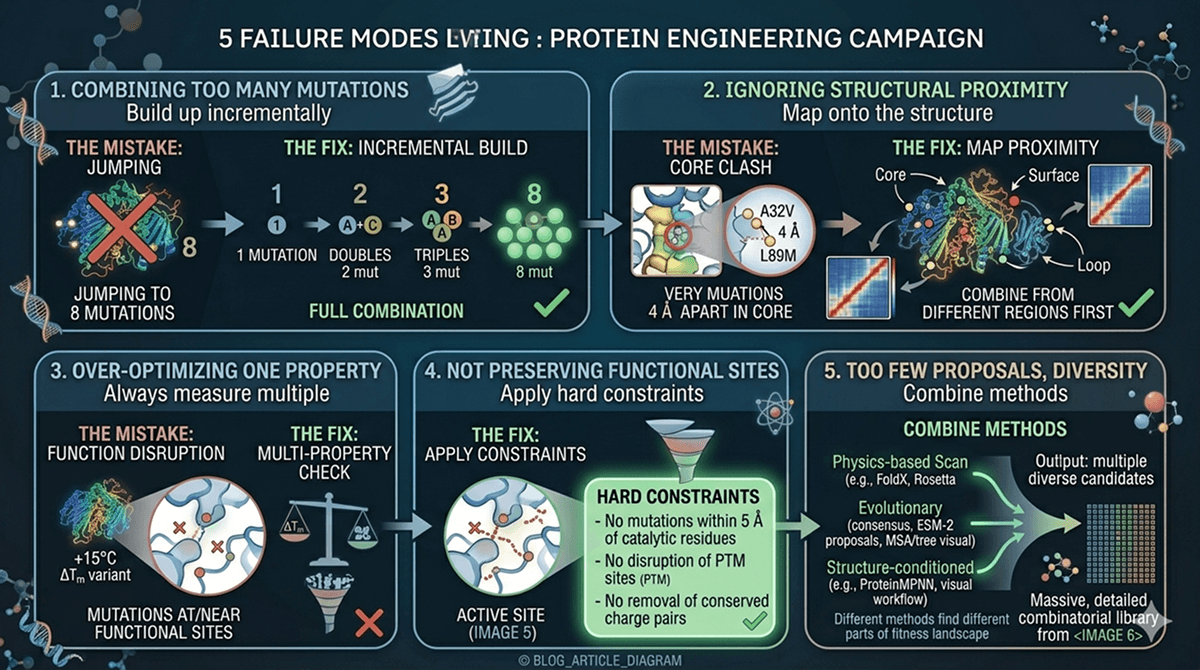
Common Failure Modes
Failure 1: Combining Too Many Mutations at Once
The mistake: Jumping from singles directly to an 8-mutation combination.
The fix: Build up incrementally. Test doubles and triples first. If pairwise combinations work, the full combination is more likely to succeed.
Failure 2: Ignoring Structural Proximity
The mistake: Combining two mutations that are 4 Å apart in the core.
The fix: Map mutations onto the structure. Mutations that are structurally close are more likely to show epistasis. Combine mutations from different regions first.
Failure 3: Over-Optimizing One Property
The mistake: Maximizing ΔTm without checking activity, expression, or aggregation.
The fix: Always measure multiple properties. A +15°C ΔTm variant with 10% residual activity is useless for most applications.
Failure 4: Not Preserving Functional Sites
The mistake: Allowing mutations at or near active sites, binding sites, or PTM sites.
The fix: Apply hard constraints: no mutations within 5 Å of catalytic residues, no disruption of known PTM sites, no removal of conserved charge pairs.
Failure 5: Too Few Proposals, Too Little Diversity
The mistake: Relying on a single proposal method (e.g., only Rosetta).
The fix: Combine physics-based, evolutionary, and structure-conditioned proposals. Different methods find different parts of the fitness landscape.
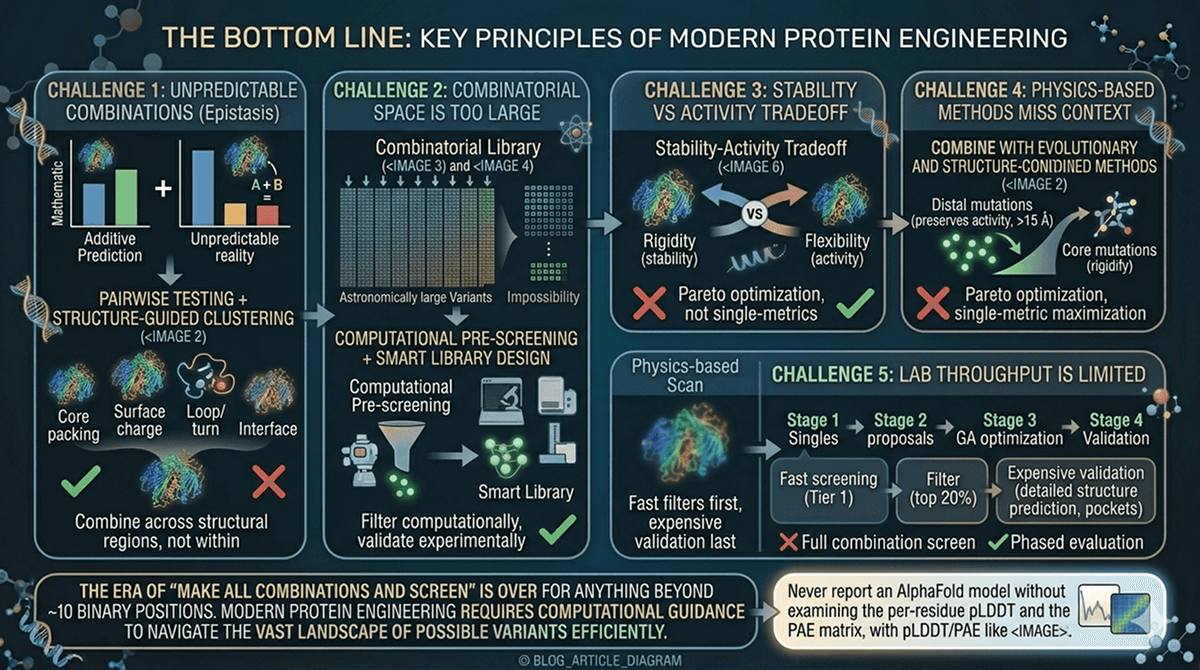
The Bottom Line
Challenge | Solution | Key Principle |
|---|---|---|
Epistasis makes combinations unpredictable | Pairwise testing + structure-guided clustering | Combine across structural regions, not within |
Combinatorial space is too large | Computational pre-screening + smart library design | Filter computationally, validate experimentally |
Stability vs activity tradeoff | Multi-objective optimization, distal mutations | Pareto optimization, not single-metric maximization |
Physics-based methods miss context | Combine with evolutionary and structure-conditioned methods | Diverse proposals find diverse solutions |
Lab throughput is limited | Staged evaluation with increasing stringency | Fast filters first, expensive validation last |
The era of "make all combinations and screen" is over for anything beyond ~10 binary positions. Modern protein engineering requires computational guidance to navigate the vast landscape of possible variants efficiently.
Scaling Variant Optimization with Orbion
For researchers navigating the singles-to-combinations challenge, Orbion's Mutation Engine implements the multi-stage genetic algorithm approach described above. Stage 1 performs a physics-based scan (hydrophobic packing, conservative substitutions, aromatic clusters). Stage 2 diversifies with ESM-2 and ProteinMPNN proposals. Stage 3 runs combinatorial optimization seeded from top singles. Stage 4 exports candidates for analysis.
The Stabilize module scores any variant—single or multi-mutation (e.g., A123G/K456R)—with AstraDTM (ΔTm) and AstraDDG (ΔΔG), along with biophysical metrics like Δdisorder and Δamyloidogenicity. Batch CSV import handles up to 100 variants per upload, and top candidates can be validated with AlphaFold2 structure prediction. Hard constraints preserve PTM sites, binding sites, and charge balance throughout the optimization—so you can explore the combinatorial landscape without accidentally breaking what matters.
References
Starr TN & Thornton JW. (2016). Epistasis in protein evolution. Protein Science, 25(7):1204-1218. PMC4721762
Romero PA & Arnold FH. (2009). Exploring protein fitness landscapes by directed evolution. Nature Reviews Molecular Cell Biology, 10:866-876. Link
Kellogg EH, et al. (2011). Role of conformational sampling in computing mutation-induced changes in protein structure and stability. Proteins, 79(3):830-838. PMC3076786
Schymkowitz J, et al. (2005). The FoldX web server: an online force field. Nucleic Acids Research, 33:W382-W388. PMC1325220
Dauparas J, et al. (2022). Robust deep learning-based protein sequence design using ProteinMPNN. Science, 378(6615):49-56. Link
Meier J, et al. (2021). Language models enable zero-shot prediction of the effects of mutations on protein function. NeurIPS, 35. PMC8886682
Gaucher EA, et al. (2008). Palaeotemperature trend for Precambrian life inferred from resurrected proteins. Nature, 451:704-707. Link
Risso VA, et al. (2013). Hyperstability and substrate promiscuity in laboratory resurrections of Precambrian β-lactamases. Journal of the American Chemical Society, 135(8):2899-2902. PMC3660565
Goldenzweig A, et al. (2016). Automated structure- and sequence-based design of proteins for high bacterial expression and stability. Molecular Cell, 63(2):337-346. PMC4023753
Eriksson AE, et al. (1992). Response of a protein structure to cavity-creating mutations and its relation to the hydrophobic effect. Science, 255(5041):178-183. PMC2242367
Matthews BW, et al. (1987). Enhanced protein thermostability from site-directed mutations that decrease the entropy of unfolding. Proceedings of the National Academy of Sciences, 84(19):6663-6667. PMC2144110
Buss O, et al. (2018). FoldX as protein engineering tool: better than random based approaches? Computational and Structural Biotechnology Journal, 16:25-33. PMC6820749
Makowski EK, et al. (2021). Co-optimization of therapeutic antibody affinity and specificity using machine learning models that generalize to novel mutational space. Nature Communications, 13:3788. PMC7655765
